Extending the shelf life of antibody drugs
A new computer model developed at MIT can help solve a problem that has plagued drug companies trying to develop promising new treatments made of antibodies: Such drugs have a relatively short shelf life because they tend to clump together, rendering them ineffective.
Antibodies are the most rapidly growing class of human drugs, with the potential to treat cancer, arthritis and other chronic inflammatory and infectious diseases. About 200 such drugs are now in clinical trials, and a few are already on the market.
Patients can administer these drugs to themselves, but this requires high doses — and the drugs must therefore be stored at high concentrations. However, under these conditions the drugs tend to clump, or aggregate. Even if they are stored at lower concentrations and administered by a doctor intravenously, they often have stability issues. Addressing such issues typically takes place later in the drug development process, and the cost — both in time and money — is often high.
Currently there is no straightforward way to address these storage issues early in the development process.
"Drugs are usually developed with the criteria of how effective they'll be, and how well they'll bind to whatever target they're supposed to bind," says Bernhardt Trout, professor of chemical engineering and leader of the MIT team. "The problem is there are all of these issues down the line that were never taken into account."
Trout and his colleagues, including Bernhard Helk of Novartis, have developed a computer model that can help designers identify which parts of an antibody are most likely to attract other molecules, allowing them to alter the antibodies to prevent such clumping. The model, which the researchers aim to incorporate in the drug discovery process, is described in a paper appearing in the online edition of the Proceedings of the National Academy of Sciences the week of June 29.
Preventing aggregation
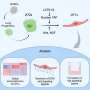
Most of the aggregation seen in antibodies is due to interactions between exposed hydrophobic (water-fearing) regions of the proteins.
Trout's new model, known as SAP (spatial aggregation propensity), offers a dynamic, three-dimensional simulation of antibody molecules. Unlike static representations such as those provided by X-ray crystallography, the new model can reveal hydrophobic regions and also indicates how much those regions are exposed when the molecule is in solution. The other important aspect of the model is that it selects out regions responsible for aggregation, as opposed to just single sites.
Once the hydrophobic regions are known, researchers can mutate the amino acids in those regions to decrease hydrophobicity and make the molecule more stable. Using the model, the team produced mutated antibodies with greatly enhanced stability (up to 50 percent more than the original antibodies), and the mutations had no adverse affect on their function.
Source: Massachusetts Institute of Technology