Scientists find two compounds that lay the foundation for a new class of AIDS drug
A team of scientists at The Scripps Research Institute has identified two compounds that act on novel binding sites for an enzyme used by the human immunodeficiency virus (HIV), the virus that causes AIDS. The discovery lays the foundation for the development of a new class of anti-HIV drugs to enhance existing therapies, treat drug-resistant strains of the disease, and slow the evolution of drug resistance in the virus.
The research will appear as the cover story of the March issue of the journal Chemical Biology & Drug Design.
The anti-HIV compounds identified in the new study bind to HIV protease—an enzyme essential to the lifecycle of the virus. Drugs that block this viral enzyme are called "protease inhibitors" and currently make up an important part of the successful AIDS drug cocktail known as highly active anti-retroviral therapy (HAART).
Compared with the nine U.S. Food and Drug Administration (FDA)-approved drugs that target HIV protease, however, the two new compounds, which are small chemical units or "fragments," bind with two novel parts of the molecule. This could make future drugs incorporating the fragments' novel structural elements a useful complement to existing treatments.
"The study's results open the door to a whole new approach to drug design against HIV protease," said Scripps Research Associate Professor C. David Stout, senior author of the study. "The fragments bound at not one, but two, different crevices in protease outside the active site. This is an important proof-of-concept that the protease molecule has two non-active site binding pockets ('allosteric sites') which can now be exploited as a powerful new strategy to combat drug-resistance in HIV."
Research Associate Alex L. Perryman, first author of the paper and member of Professor Arthur Olson's laboratory at Scripps Research, added, "The experiments validate my hypothesis developed from computational modeling that HIV protease has pockets on its surface besides the active site that can bind drugs. Drugs developed to target these sites could be used to make current FDA-approved active site inhibitors more potent and to restore their effectiveness against drug-resistant superbugs. The whole strategy of targeting non-active sites may also prove useful against other diseases, especially when there are mutations that cause drug resistance."
The Long Arms of HIV Protease
Approximately 33 million people are currently living with HIV infections, 2.7 million of whom were newly infected in 2008, according to the most recent statistics available from the World Health Organization. While HAART therapy has prolonged survival and improved the quality of life for many AIDS patients, a rapidly growing portion of new HIV infections involve drug-resistant strains.
Perryman has been interested in the problem of drug resistance for some time. As a Howard Hughes Medical Institute (HHMI) pre-doctoral fellow in the laboratory of HHMI Professor J. Andrew McCammon at the University of California, San Diego, Perryman conducted extensive computer simulations of a drug-resistant HIV protease molecule and compared them with a normal HIV protease molecule. This theoretical effort laid the groundwork for the current experimental study.
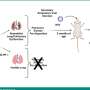
As is often the case in biology, the shape of the HIV protease reflects and dictates its function. In the process of reproducing itself, the virus makes long protein chains, which the protease splits into shorter pieces to enable the final assembly of new HIV virions. To split off the pieces from the longer chain, the protease has two scissor-like flaps on each side of the molecule. The flaps open to let in the viral protein chain, which threads itself into the active site binding pocket. The flaps then close, crossing in the middle over a binding pocket and cleaving the protein chain into smaller pieces.
Current HIV-protease drugs mimic the shape of certain regions of the HIV protein chain (i.e., the "cleavage sites") and bind to the active site in the hollow center of protease. Once this site is blocked by the drug, the protease is disabled, and HIV cannot make new infectious particles. Thus, protease drugs impede the spread of the HIV infection to other cells within a patient.
Perryman wanted to find out what was different about drug-resistant strains of the virus—information that was not obvious from static pictures of the molecules. So, he conducted computer simulations of the movement of a particularly nasty multi-drug-resistant mutant strain of HIV (V82F/I84V).
The findings showed that the flaps of the drug-resistant protease molecule tended to be open more often than their standard counterparts, and they were also more flexible. While the anti-HIV drugs still fit into the active site binding pocket, more energy was needed to close the flaps than the drugs could muster. As a consequence, the drugs wouldn't stay in the binding site, and the pocket remained available for the HIV protein chain, which was still able to close the flaps and go on to create new infectious particles.
In their simulations, Perryman and his colleagues identified a potential solution to this problem. Like restraining the handle-end of a pair of scissors to keep the blades from opening too wide, a new type of drug might be able to bind to alternate sites on the sides of the protease, restraining the flaps from their ends and providing the current anti-HIV drugs enough help to close the flaps and disable the protease. Instead of blocking the protease's active site, these compounds would be "allosteric fragments," small molecule building blocks that shift the dynamics of the molecule so it prefers a different conformation (shape).
The plan sounded good in theory, but could it work in practice?
Putting Together the Pieces
That's what the Scripps Research team set out to learn in the new study, which was funded by a grant to the Olson lab from the National Institutes of Health to seek ways to combat drug-resistance in HIV.
To accomplish this goal, the team procured a "library" of 384 compound fragments that had been compiled by a company called Active Sight. Screening fragments rather than larger compounds has become more popular over the last several years, Perryman explained, since using these smaller pieces is a more efficient way to find promising structures than using libraries of large compounds. In addition, fragments can be extended, combined, and modified using "structure-based drug design" in a way that makes them fit tightly into the right binding sites, without displaying the unwanted interactions that sometimes come with larger molecules. Using libraries of much larger, "lead-like" compounds is less efficient, and those large compounds are less extendable.
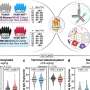
Since the HIV protease molecule is in constant motion, the team of scientists, which included members of Scripps Research Professor John Elder's lab, first crystallized the molecule in various different conformations. The scientists then screened a library of fragments against these crystals to see if any of them bound and characterized the structural features of the results using the Stanford Synchrotron Radiation Lightsource (SSRL). Stout notes that, in total, the project conducted a massive screen of more than 800 crystals producing more than 400 data sets, a feat made possible by SSRL's robotic capabilities.
In their initial experiments, the scientists met with partial success—enough to establish a proof-of-concept—as one fragment attached to the "eye site" between the tip of a flap and the top of the active site's wall. However, the large active site of the molecule tended to be problematic, interfering with the scientists' goal of searching for fragments that bind to alternate sites on the molecule.
To overcome this problem, the team "plugged up" the active site with a known inhibitor so that the screen would identify only fragments that bound to other regions of the protease. Perryman noted that this innovative protocol could be applied to similar work on many different disease-causing agents, especially those that have developed drug resistance.
Using the new method for crystallographic screening, the team found two "hits"—fragments 2-methylcyclohexanol and indole-6-carboxylic acid. The scientists used additional x-ray crystallographic experiments to confirm first that the fragments indeed bound to novel sites in the protease and second that these fragments change the structural preferences of protease.
"Since these fragments are very small, we wouldn't expect them to be potent inhibitors by themselves," said Perryman. "But it's the beginning of a process where we can try to use these little fragments to build up to a real potent inhibitor. The study validates our predictions and lays the structural foundation for the development of new classes of anti-AIDS drugs."
More information: www3.interscience.wiley.com/jo … l/123244298/abstract