Researchers discover genes involved in colorectal cancer
A jumping gene with the fairy tale name "Sleeping Beauty" has helped to unlock vital clues for researchers investigating the genetics of colorectal cancer.
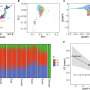
A study published today used the Sleeping Beauty transposon system to profile the repertoire of genes that can drive colorectal cancer, identifying many more than previously thought. Around one third of these genes are mutated in human cancer, which provides strong evidence that they are driver mutations in human tumours.
The collaborative project funded by Cancer Research UK and the Wellcome Trust was led by Dr David Adams from the Wellcome Trust Sanger Institute, and Dr Douglas Winton, of the Cancer Research-UK Cambridge Research Institute.
"These findings, when combined with mutation data from human colon cancers, will drive forward our understanding of the processes that lead to colorectal cancer," says Dr Adams, senior author from the Sanger Institute. "They demonstrate how many genes can contribute to this cancer and how these genes work together in the development of this disease".
The Sleeping Beauty transposon system induces genetic mutations at random, identifying and tagging candidate cancer genes, the drivers that cause colorectal cancer. This system has become critical in uncovering the genetic pathways that cause cancer, and, in this study, the team identify more than 200 genes that can be disrupted in human colorectal cancers.
Colorectal (bowel) cancer is the third most common cancer in the UK, and the second most common cause of cancer deaths after lung cancer; just under 40,000 people were diagnosed with bowel cancer in the UK in 2008 – around 110 people every day – a figure which has shown little improvement over the last decade.
"Our research provides a rich source of candidate genes that represent potential diagnostic, prognostic and therapeutic targets, and defines the breadth of genes that can contribute to cancer of the intestine," says Dr Winton, senior author from the Cancer Research UK Cambridge Research Institute. "It is becoming increasingly clear that cancers are driven by mutations in disparate collections of genes and it is essential that we tease apart the important changes."
Current thinking is that perhaps around 50 major drivers are mutated in any one cancer cell, but the number and identity of all of the cancer drivers, and how many drivers are found in each type of cancer, is largely unknown. By performing screens for cancer genes in the mouse and by then comparing them to data from human tumours the team identified a rich catalogue of new candidate genes helping to refine the genes that genetic pathways that drive bowel cancer development.
"At its heart, cancer is a disease driven by faulty genes," says Dr Lesley Walker, director of cancer information at Cancer Research UK. "Research suggests that each cancer cell has a number of 'driver' faults that make them grow out of control, as well as 'passenger' faults that they pick up as the disease develops. This technique is helping us to tease out the key drivers of bowel cancer, laying the foundations for more effective, targeted treatments for the disease in the future."
The research complements studies by The Cancer Genome Atlas and the International Cancer Genome Consortium, which are cataloguing the mutations responsible for cancer development using next generation DNA sequencing.
More information: Nature Genetics doi: 10.1038/ng.990