Scientists provide global view of how HIV hijacks cells during infection
In perhaps the most comprehensive survey of the inner workings of HIV, an international team of scientists led by researchers at the University of California, San Francisco has mapped every apparent physical interaction the virus makes with components of the human cells it infects—work that may reveal new ways to design future HIV/AIDS drugs.
Explored this week in back-to-back papers in the journal Nature, the survey reveals a pathogenic landscape in which HIV's handful of proteins makes hundreds of physical connections with human proteins and other components inside the cell.
In one paper, the team details 497 such connections, only a handful of which had been previously recognized by scientists. Disrupting these connections may interfere with HIV's lifecycle, and the existence of so many new connections suggests there may be several novel ways to target the virus.
"Have we identified new drug targets?" said Nevan Krogan, PhD, who led the research. "I believe we have."
Krogan is an associate professor of Cellular and Molecular Pharmacology at UCSF and an affiliate of the California Institute for Quantitative Biosciences (QB3). He is also a faculty member at the UCSF‑affiliated J. David Gladstone Institutes in San Francisco. Stefanie Jager, the lead author from the Krogan lab, collaborated with several other labs in the UCSF School of Medicine and the UCSF School of Pharmacy ‑ including those of Andrej Sali, Alan Frankel, Charles Craik, Alma Burlingame, Ryan Hernandez and Tanja Kortemme ‑ to carry out the global analysis.
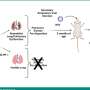
In a companion paper, Krogan and collaborating labs of John Gross (UCSF) and Reuben Harris (University of Minnesota) investigated one such connection in detail. They discovered that an HIV protein called Vif makes a physical connection with a human protein called CBF-β, hijacking its function. This virus requires this action to function, said Krogan, which suggests that disrupting the connection may be a viable way to design new HIV/AIDS therapies.
"This is a good example of how biophysical studies can improve our understanding of disease and point the way to the exploration of potential therapeutic targets," said Judith H. Greenberg, PhD, acting director of the National Institute of General Medical Sciences (NIGMS).
NIGMS and the National Institute of Allergy and Infectious Diseases (NIAID), both components of the National Institutes of Health, partially funded this work through three large structural and systems biology program grants to UCSF—called HARC (HIV Accessory and Regulatory Complexes) (PI: Alan Frankel, UCSF), HINT (HIV Innate Networks Team) (PI: John Young, Salk Institute) and the Center for Systems and Synthetic Biology (PI: Wendell Lim, UCSF). The work was also funded by NIH's National Center for Research Resources, QB3 and through a Searle Scholarship and a Keck Young Investigator Award. Several of the researchers are also associated with the Host Pathogen Circuitry Center at UCSF.
"These papers represent a truly collaborative effort from many different groups from several universities," Krogan said, "The multi-disciplinary nature of this work is key if we are to tackle problems of this magnitude."
Unlocking the Doors to HIV Drug Design
Understanding the details of how HIV works within human cells and finding ways to interfere with those processes are two of the central questions scientists start with when designing drugs to treat people with HIV/AIDS.
The virus is small and has only a tiny amount of genetic material in its own genome—about 10,000 times less DNA than that of the human genome. Where typical human cells may include thousands of different proteins, HIV brings fewer than 20 of its own. Once inside a human cell, however, the virus relies extensively on the existing molecular machinery found there, hijacking the function of some human proteins, shutting down others and subverting still more.
HIV is remarkably effective at this. In the last 30 years, the virus has infected tens of millions worldwide. The World Health Organization estimates some 34 million people are currently living with the virus and that HIV has already claimed some 25 million lives.
One of the greatest success stories in the last few decades has been the development of powerful antiretroviral drugs that block the action of various viral proteins essential to HIV's lifecycle. Since the 1990s, the standard of treatment for people with HIV/AIDS has been therapy that combines several of these drugs. The availability of such drug combinations worldwide has allowed millions of people with the disease to live longer.
The UCSF-led study has provided the most comprehensive and detailed picture to date of all the interactions HIV has with the human cells it infects, and identifying these interactions may lead to the development of new drugs to treat the disease.
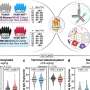
Of the 497 specific interactions between HIV and human proteins discovered in the new work, only 19 of those were previously reported. What accounts for the discrepancy, Krogan said, is that this was the first study to look for such interactions globally and in an unbiased fashion – unlike previous studies, which had been more focused. The work involved expressing every HIV protein in a series of human cells, pulling these viral proteins out of the cells, analyzing them with a method known as mass spectrometry and looking for the associated human proteins.
To illustrate the potential significance of these new associations, Krogan and his colleagues explored one of them in depth in the second paper: an association between an HIV protein called Vif and CBF-β, a human protein.
Vif is known to target for degradation the restriction factor APOBEC3G, a host factor that protects cells from HIV infection. Krogan and colleagues show that the transcription factor CBF-β associates with Vif and is needed for Vif to counteract APOBEC3G. Previously no one had known CBF-β collaborated with Vif to play such a role, Krogan said, and interfering with this association may be a way to block the virus. Ultimately, if scientists can design compounds to do this safely and effectively, those compounds could form the basis for a new type of HIV/AIDS treatment.
More information: The article, "Global landscape of HIV–human protein complexes" by Stefanie Jäger, Peter Cimermancic, Natali Gulbahce, Jeffrey R. Johnson, Kathryn McGovern, Starlynn C. Clarke, Michael Shales, Gaelle Mercenne, Lars Pache, Kathy Li, Hilda Hernandez, Gwendolyn M. Jang, Shoshannah L. Roth, Eyal Akiva, John Marlett, Melanie Stephens, Ivan D'Orso, Jason Fernandes, Marie Fahey, Cathal Mahon, Anthony J. O'Donoghue, Aleksandar Todorovic, John H. Morris, David A. Maltby, Tom Alber, Gerard Cagney, Fredric D. Bushman, John A. Young, Sumit K. Chanda, Wesley I. Sundquist, Tanja Kortemme, Ryan D. Hernandez, Charles S. Craik, Alma Burlingame, Andrej Sali, Alan D. Frankel and Nevan J. Krogan is published online on December 21, 2011. dx.doi.org/10.1038/nature10719
The article, "Vif hijacks CBF-b to degrade APOBEC3G and promote HIV-1 infection" by Stefanie Jäger, Dong Young Kim, Judd F. Hultquist, Keisuke Shindo, Rebecca S. LaRue, Eunju Kwon, Ming Li, Brett D. Anderson, Linda Yen, David Stanley, Cathal Mahon, Joshua Kane, Kathy Franks-Skiba, Peter Cimermancic, Alma Burlingame, Andrej Sali, Charles Craik, Reuben S. Harris, John D. Gross and Nevan J. Krogan is published online on December 21, 2011. dx.doi.org/10.1038/nature10693