The genetic basis for age-related macular degeneration
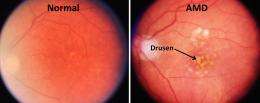
Age-related macular degeneration (AMD) is one of the leading causes of blindness worldwide, especially in developed countries, and there is currently no known treatment or cure or for the vast majority of AMD patients. New research published in BioMed Central's open access journal Genome Medicine has identified genes whose expression levels can identify people with AMD, as well as tell apart AMD subtypes.
It is estimated that 6.5% of people over age 40 in the US currently have AMD. There is an inheritable genetic risk factor but risk is also increased for smokers and with exposure to UV light. Genome-wide studies have indicated that genes involved in the innate immune system and fat metabolism are involved in this disease. However none of these prior studies examined gene expression differences between AMD and normal eyes.
In order to address this question, researchers at the University of California Santa Barbara, the University of Utah John Moran Eye Center, and the University of Iowa combined forces and used a human donor eye repository to identify genes up-regulated in AMD. The ability of these genes to recognize AMD was tested on a separate set of samples.
The team discovered over 50 genes that have higher than normal levels in AMD, the top 20 of which were able to 'predict' a clinical AMD diagnosis. Genes over-expressed in the RPE-choroid (a tissue complex located beneath the retina) included components of inflammatory responses, while in the retina, the researchers found genes involved in wound healing and the complement cascade, a part of the innate immune system. They found retinal genes with expression levels that matched the disease severity for advanced stages of AMD.
Dr. Monte Radeke, one of the project leaders, explained, "Not only are these genes able to identify people with clinically recognized AMD and distinguish between different advanced types – some of these genes appear to be associated with pre-clinical stages of AMD. This suggests that they may be involved in key processes that drive the disease. Now that we know the identity and function of many of the genes involved in the disease, we can start to look among them to develop new diagnostic methods, and for new targets for the development of treatments for all forms of AMD."
More information: Systems-level analysis of age-related macular degeneration reveals global biomarkers and phenotype-specific functional networks, Aaron M Newman, Natasha B Gallo, Lisa S Hancox, Norma J Miller, Carolyn M Radeke, Michelle A Maloney, James B Cooper, Gregory S Hageman, Don H Anderson, Lincoln V Johnson and Monte J Radeke, Genome Medicine (in press) genomemedicine.com/