New research may have discovered how memories are encoded in our brains
University of Alberta led research may have discovered how memories are encoded in our brains.
Scientists understand memory to exist as strengthened synaptic connections among neurons. However components of synaptic membranes are relatively short-lived and frequently re-cycled while memories can last a lifetime.
Based on this information, U of A physicist and lead researcher Jack Tuszynski, his graduate student Travis Craddock and University of Arizona professor Stuart Hameroff investigated the molecular mechanism of memory encoding in neurons.
The team looked into structures at the cytoskeletal level of brain structure. They found components that fit together and were capable of creating the information processing and storage capacity that the brain needs to form and retain memory.
The practical implications of understanding the mechanism of memory encoding are enormous.
"This could open up amazing new possibilities of dealing with memory loss problems, interfacing our brains with hybrid devices to augment and 'refresh' our memories," says Tuszynski. "More importantly, it could lead to new therapeutic and preventive ways of dealing with neurological diseases such as Alzheimer's and dementia, whose incidence is growing very rapidly these days."
Q & A with Jack Tuszynski
Why did you decide to tackle the question of where the information pertinent to memory is stored?
This is one of the key open and most intriguing problems in neuroscience. We read a paper on experiments that were able to safely erase memories in animals. These experiments implicated a specific molecule (the enzyme called calcium-calmodulin dependent kinase complex II) that is instrumental in encoding and erasing memory in the brain. We noticed an amazing similarity in its geometry to a patch of tubulin dimers in a microtubule. Microtubules fill the interiors of our brain's neurons, especially in axons and dendrites where most of the activity takes place. We undertook to find out if this similarity is accidental or not. This led to the generation of a very accurate computational model of the interaction between CaMKII and microtubules. It looks to us that a mechanism for memory encoding has now been found.
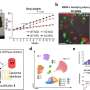
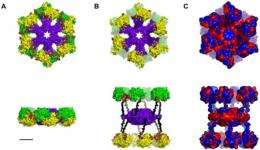
What are we looking at in figures 1 through 6 of the paper?
These figures were generated by a computational reconstruction of the molecules that play key roles in memory encoding: tubulin, microtubules and CaMKII. We know their atomic-resolution 3D structures from crystallographic data that are publicly available. We then built computer models of higher level organization of these protein complexes. These models allow us to test the spatial compatibility of the protein-protein interactions as well as to calculate interaction energies to see if they attract themselves under specific conditions and if so, how strongly.
Could an understanding of this mechanism lead to improvements or repairs to the memory function and could memories themselves be restored?
Yes, all of this is entirely possible. We can potentially use this insight to improve quality of memory storage, memory recall, information processing as well as the amount of data stored.
Are you planning follow-up research to this paper?
We are already working on various applied aspects related to this paper. Another paper has been just accepted for publication, based on our collaboration with University of Arizona, Harvard and Boston University, where we propose a molecular mechanism of Alzheimer's disease progression. This should soon be published in PLos One and it'll lead to specific therapeutic solutions as our next project. It turns out that the concentration of zinc plays a crucial role in stabilizing microtubules but also in fueling the buildup of amyloid plaques. We developed a physical model of the process. A third paper in this series stemming from Travis Craddock's PhD thesis discusses the effects of anesthetic molecules on memory blockage in brain's microtubules.
More information: Their paper, "Cytoskeletal Signaling: Is Memory Encoded in Microtuble Lattices by CaMKII Phosphorylation?" was published in the journal, PLoS Computational Biology.