Researchers search public databases, flag novel gene's key role in type 2 diabetes
Using computational methods, Stanford University School of Medicine investigators have strongly implicated a novel gene in the triggering of type-2 diabetes. Their experiments in lab mice and in human blood and tissue samples further showed that this gene not only is associated with the disease, as predicted computationally, but is also likely to play a major causal role.
In a study to be published online April 9 in Proceedings of the National Academy of Sciences, the researchers combed through freely accessible public databases storing huge troves of results from thousands of earlier experiments. They identified a gene never before linked to type-2 diabetes, a life-shortening disease that affects 4 percent of the world's population. These findings have both diagnostic and therapeutic implications.
The study's senior author is Atul Butte, MD, PhD, associate professor and chief of systems medicine in pediatrics; its first author is Keiichi Kodama, MD, PhD, a staff research scientist in Butte's group.
Ordinarily, cells throughout the body, alerted to the presence of sugar in the blood by insulin, hungrily slurp it up for use as an energy source. But excessive blood-sugar levels — diabetes' defining feature — eventually damage blood vessels, nerves and other tissues.
There are two broad categories of diabetes. In type-1 diabetes, a relatively rare autoimmune condition that typically begins in childhood, insufficient insulin is secreted by the pancreas. Type-2 diabetes, on the other hand, results from a phenomenon called insulin resistance: the tendency of cells in tissues throughout the body — but especially in fat, liver and muscle — to lose sensitivity and ignore the insulin's "gravy train" signal.
Drugs now used to treat insulin resistance can't reverse the progression to full-blown type-2 diabetes. "We don't really have a good grasp of the molecular pathology that makes people get it in the first place," said Butte.
In searching for risk-increasing genes over the past 10 years, scientists have used two approaches to hunt them down. One way is to look for variations in genes' composition — deviations in their chemical sequences that correlate with a higher likelihood of contracting a particular condition.
But genes don't change from one tissue to the next, and — with the exception of mutations that accrue gradually over a lifetime in particular cells and can lead to cancer and other conditions — they remain largely unaltered by disease and the aging process. What does change dynamically, from one tissue or state to another, is what all those genes are doing: how actively each of them is involved in cranking out the starting materials for the many thousands of proteins critical to each cell's or tissue's identity and to every organism's survival. In any given cell in a person's body, at any given time, some genes are switched off, others somewhat on and still others working overtime.
And so a second approach to understanding our genes has been devised. This latter method flags differences in genes' activity levels, for example in diseased vs. normal tissues, for each of the 20,000 genes in the entire genome.
Both types of approaches have generated staggering amounts of data — far more than can fit onto the pages of standard, peer-reviewed journals, whose editors routinely demand (as do federal-government funding agencies) that researchers park their experiments' results in online, public repositories accessible to others. Now, investigators such as Butte are starting to reach in, drill down and pull out a treasure-trove of potentially valuable information.
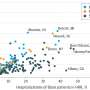
In this study, the Stanford scientists wanted to know which genes showed especially marked changes in activity, as indicated in earlier comparisons of diabetic vs. healthy tissue samples (notably fat, muscle, liver and beta cells, the only cells in the body that release insulin). Mining public databases, they located 130 independent gene-activity-level experiments — in rats, mice and humans — comprising 1,175 separate individual samples in all. Then, integrating that data, they searched for those genes that showed activity-level differences in the most experiments.
They zeroed in on a single gene, called CD44, whose activity changed substantially in diabetic tissues compared with healthy tissues in 78 of the 130 experiments. The chance of this occurring "just due to dumb luck," Butte said, was vanishingly small: less than one in 10 million-trillion. The uptick in CD44's activity was especially pronounced in the fat tissue of people with diabetes, he said — intriguing, because obesity is known to be a strong risk factor for type-2 diabetes.
The gene was interesting in itself. CD44 codes for a cell-surface receptor not found on fat cells, although those cells do have surface molecules that bind to it. Rather, this receptor sits on the surface of scavenger cells called macrophages (from the Greek words for "big eater") that can cause inflammation. In obese individuals, macrophages migrate to and take up positions in fat tissue. (Indeed, as many as half the cells in a big potbelly can be macrophages.) Recent medical research has strongly implicated inflammation in initiating type-2 diabetes.
CD44 was first identified more than a decade ago by immunologists looking for a possible connection to autoimmune disease. To test that connection, those immunologists created a strain of laboratory mouse lacking the gene. By chance, these "CD44 knockout" mice were derived from a lab-mouse strain that, if fed a high-fat diet, has a propensity for becoming obese, insulin-resistant and diabetic. With the exception of that missing gene, these two strains are identical.
Butte and his colleagues obtained these two strains of mice (one carrying the gene, the other lacking it) and divided them into two subgroups, which they fed either a normal or a high-fat diet, representative of today's increasingly common human diet. The team studied these mice using tests commonly applied to humans, for example measuring fasting blood sugar and measuring blood sugar after administering sugar or insulin. As anticipated, the mice on normal diets stayed slim and retained good insulin sensitivity. CD44-containing mice on high-fat diets, also as expected, got tubby and developed insulin resistance. But mice lacking the suspect gene never lost their sensitivity to insulin and didn't become diabetic on high-fat diets, although they became as plump as their CD44-carrying peers.
This suggested that knocking out CD44's function could improve insulin sensitivity, and that blocking CD44 with a drug might turn out to be an interesting new way to treat type-2 diabetes. So the team tested a prototype drug: antibodies that shut down the receptor's action in CD44-carrying mice fed a high-fat diet. Though these overfed mice didn't get any thinner, the prototype drug did reduce their blood-sugar levels within a week. Moreover, the number of macrophages in these mice's fat tissue plummeted.
Turning to human blood samples, Butte and his associates found that insulin-resistant people (those prone to developing type-2 diabetes) have higher levels of free-roving CD44-receptor molecules circulating in their blood than do people with normal insulin-processing capability. This suggests a potential early diagnostic test, or biomarker, that could help detect or predict insulin resistance. Plus, the antibody results suggest, a small molecule that blocked this receptor could have profound therapeutic potential for type-2 diabetes.