Researchers present new findings for glioblastoma at American Association for Cancer Research
Physician-scientists from University Hospitals (UH) Case Medical Center's Seidman Cancer Center and Case Western Reserve University School of Medicine presented new research findings in 24 presentations this week at the Annual Meeting of the American Association for Cancer Research (AACR) in Chicago, Illinois.
"The breadth and depth of this innovative cancer research presented at AACR is truly outstanding," says Stan Gerson, MD, Director of the Seidman Cancer Center at UH Case Medical Center and the Case Comprehensive Cancer Center at Case Western Reserve University. "Our faculty members are making tremendous advances in hematology and oncology which is reflected in their being chosen for oral and poster presentations."
Two innovative studies are investigating novel methods that may help clinicians bring a greater specificity to the treatment of glioblastoma (GBM) in the future. GBM is the most common malignant primary brain tumor in adults and also the most aggressive. Median survival time for GBM is approximately 12-15 months.
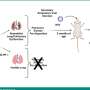
A study led by Jill Barnholtz-Sloan, PhD, Associate Professor at Case Comprehensive Cancer Center and Case Western Reserve University School of Medicine, sought to identify protein biomarkers that can help physicians determine which patients may benefit from standard treatment for GBM. Using research tumor samples from patients with short-term survival (defined as < nine months after diagnosis) and patients with long-term survival (who lived greater than 18 months after diagnosis), the investigators found 183 proteins to be significant between survival groups. Biomarkers have been identified in other cancers such as breast and colon but progress in treatment for GBM patients has been slowed by the absence of biomarkers to define treatment response.
"This has the potential to enable physicians to tailor individualized treatments for specific brain cancer patients," said Andrew Sloan, MD, Co-Investigator on the study and Director of the Brain Tumor and Neuro-Oncology Center at UH Case Medical Center and Associate Professor at Case Western Reserve University School of Medicine. "Current standards are to treat every patient in the same fashion but if we can identify who will benefit from specific therapies, this represents an important step forward for brain tumor treatment."
The researchers are also validating these results against clinical tumor samples from another group of patients. In addition to verifying their results in this way, Dr. Barnholtz-Sloan said the hope is to do a larger study involving all the hospitals in the Ohio Brain Tumor Study, a multi-site consortium within Ohio, and to eventually develop a test for patients that could be used in a typical hospital clinical setting in order to select patients for either standard treatment or an alternative treatment.
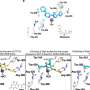
The second study, led by Vishal Patel, a MD/PHD student at Case Western Reserve University School of Medicine, in collaboration with Drs. Sloan and Barnholtz-Sloan and Dr. Mark Chance, Vice Dean for Research and Director of the Proteomics and Bioinformatics Center at the Case Western Reserve University School of Medicine, utilized a "systems" approach to finding networks of genes that may characterize short-term GBM survivors from long-term GBM survivors, rather than looking for individual proteins.
"The systems approach looks at a 'symphony' of proteins, rather than the 'solo players' that the first study used," said Dr. Sloan. "We looked at groups of related proteins connected to patient survival".
In the study, the investigators found that eight network targets were significantly differentially expressed between short-term and long-term survivors, and expression levels of three of the proteins predicted short term versus long term survival with 75 percent or more accuracy.
The studies were funded by National Cancer Institute grants to Case Western Reserve University as well as Case Comprehensive Cancer Center pilot funds and the Center for Proteomics and Bioinformatics at Case Western Reserve University.