Native digestive tract bacteria help fend off invaders, study finds
From tiny villages in developing nations to suburban kitchens in the United States, dangerous strains of E. coli bacteria sicken millions of people each year – and kill untold numbers of children.
Now, new research from the University of Michigan Health System gives scientists a better understanding of what is going on in the diarrhea-wracked guts of its victims, and what might be done to prevent or treat it.
Specifically, they show that the bacteria that usually live in our digestive tracts compete against invading bacteria such as E. coli to help our bodies fend them off.
They also show that the invaders depend on certain genes to gain a temporary upper hand in that battle -- just long enough to reproduce and cause the symptoms that expel their offspring from the body so they can find a new host.
The findings, published in journal Science on its Science Express website, point to potential ways to prevent or treat infections by enterohemorrhagic or enteropathogenic E. coli. Those are the types that can lurk in undercooked ground beef, unpasteurized milk, untreated drinking water, and contaminated produce – and that can cause diarrhea and other symptoms that sicken adults and can kill vulnerable children.
"More than 1,000 species of bacteria live in our guts, in a symbiotic population called the microbiota," says Gabriel Nunez, M.D., the U-M pathologist who led the research team. "These results show that these bacteria, also called commensals, compete with pathogens (disease-causing bacteria) in a previously unappreciated way – and that the pathogens use a specific set of genes to temporarily outcompete commensals before leaving the body. Understanding this gives us potential targets for prevention and treatment."
For instance, since the research shows that harmful bacteria compete with commensal bacteria for certain nutrients that they need to survive, selectively removing some nutrients and boosting others might help. So might a more targeted use of antibiotics when treating patients who are battling an E. coli infection.
Nunez and first author Nobuhiko Kamada, Ph.D., a postdoctoral fellow, made the findings by studying mice that they infected with C. rodentium – the rodent equivalent of harmful E. coli. The study included specially bred germ-free mice that lacked all the "good" gut bacteria that normal mice and humans harbor.
Both Nunez and Kamada are members of the U-M Medical School's Department of Pathology and the U-M Comprehensive Cancer Center, and the work fits into their broader investigations of how inflammation and immunity play a role in the body's response to cancer as well as infections.
Fittingly, Nunez holds the Paul H. de Kruif Professorship in Pathology, named for the U-M graduate who wrote Microbe Hunters, a pivotal 1926 book on the history of infectious disease research.
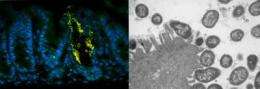
In the new paper, the team adds a new chapter to the understanding of how pathogenic bacteria gain a foothold in the gut – literally – by turning on virulence genes that allow them to attach to the cells that line the digestive tract.
This attaching-and-effacing activity, as it is called, allows the disease-causing bacteria to intimately adhere to the cells that line the gut, consume nutrients and reproduce, out-competing the natural gut bacteria. But this comfortable niche only lasts a few days or weeks, during which the host's gut gets more inflamed as the immune system responds to the insult. Diarrhea, sometimes containing blood that leaks from the gut lining, results.
And that, the researchers find, is when the pathogens stop expressing the virulence genes that allowed them to gain their upper hand. They unhitch from the gut lining, mixing in with the commensal bacteria in the open center (lumen) of the gut, and fighting for what food they can find.
While this return to competition means that some of them die, enough of them survive to be expelled in the feces. And if good sanitation systems aren't in place, the bacterial offspring have a good chance of finding a new host to take a toll on.
Better sanitation throughout the world can prevent infections in the first place, says Nunez. But when infection by pathogenic bacteria occurs, a better understanding of the way they interact with our native bacteria could eventually help save lives.
Nunez's team is working with the lab of U-M microbiologist and co-author Eric Martens, Ph.D., to screen different sugars that, if withheld or enhanced in the diet, might weaken the pathogens' effects. That could lead to a better understanding of how children and weak adults in developing nations should be fed while being treated for infection.
The University of Michigan has applied for patent protection, and is in the process of looking for commercialization partners to help bring the technology to market.
More information: Regulated Virulence Controls the Ability of a Pathogen to Compete with the Gut Microbiota, Science Express (2012).