Study explains functional links between autism and genes
A pioneering report of genome-wide gene expression in autism spectrum disorders (ASDs) finds genetic changes that help explain why one person has an ASD and another does not. The study, published by Cell Press on June 21 in The American Journal of Human Genetics, pinpoints ASD risk factors by comparing changes in gene expression with DNA mutation data in the same individuals. This innovative approach is likely to pave the way for future personalized medicine, not just for ASD but also for any disease with a genetic component.
ASDs are a heterogeneous group of developmental conditions characterized by social deficits, difficulty communicating, and repetitive behaviors. ASDs are thought to be highly heritable, meaning that they run in families. However, the genetics of autism are complex.
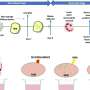
Researchers have found rare changes in the number of copies of defined genetic regions that associate with ASD. Although there are some hot-spot regions containing these alterations, very few genetic changes are exactly alike. Similarly, no two autistic people share the exact same symptoms. To discover how these genetic changes might affect gene transcription and, thus, the presentation of the disorder, Rui Luo, a graduate student in the Geschwind lab at UCLA, studied 244 families in which one child (the proband) was affected with an ASD and one was not.
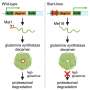
In addition to identifying several potential new regions where copy-number variants (CNVs) are associated with ASDs, Geschwind's team found genes within these regions to be significantly misregulated in ASD children compared with their unaffected siblings. "Strikingly, we observed a higher incidence of haploinsufficient genes in the rare CNVs in probands than in those of siblings, strongly indicating a functional impact of these CNVs on expression," says Geschwind. Haploinsuffiency occurs when only one copy of a gene is functional; the result is that the body cannot produce a normal amount of protein. The researchers also found a significant enrichment of misexpressed genes in neural-related pathways in ASD children. Previous research has found that these pathways include other genetic variants associated with autism, which Geschwind explains further legitimizes the present findings.
More information: Luo et al.: "Genome-wide Transcriptome Profiling Reveals the Functional Impact of Rare De Novo and Recurrent CNVs in Autism Spectrum Disorders." DOI 10.1016/j.ajhg.2012.05.011