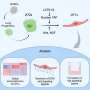
Computer model successfully predicts drug side effects
A new set of computer models has successfully predicted negative side effects in hundreds of current drugs, based on the similarity between their chemical structures and those molecules known to cause side effects, according to a paper appearing online this week in the journal Nature.
The team, co-led by researchers in the UCSF School of Pharmacy, Novartis Institutes for BioMedical Research (NIBR) and SeaChange Pharmaceuticals, Inc. – a UCSF spinoff company launched by two of the paper's authors – set out to test how well a computer model could help researchers eliminate risky drug prospects by identifying which ones were most likely to have adverse side effects.
Drugs frequently interact with more than one target, with hundreds of these targets linked to the side effects of clinically used therapeutics. Focusing on 656 drugs that are currently prescribed, with known safety records or side effects, the team was able to predict such undesirable targets – and thus potential side effects – half of the time.
That's a significant leap forward from previous work, which has never tackled hundreds of compounds at once, according to Brian Shoichet, PhD, a UCSF professor of pharmaceutical chemistry who was the joint advisor on the project alongside Laszlo Urban, MD, PhD, at Novartis.
As a result, it offers a possible new way for researchers to focus their efforts on developing the compounds that will be safest for patients, while potentially saving billions of dollars each year that goes into studying and developing drugs that fail.
"The biggest surprise was just how promiscuous the drugs were, with each drug hitting more than 10 percent of the targets, and how often the side-effect targets were unrelated to the previously known targets of the drugs," said Shoichet, whose lab is renowned for its work in using computational simulations to identify new targets for known drugs. "That would have been hard to predict using standard scientific approaches."
Adverse drug effects are the second most common reason, behind effectiveness, that potential drugs fail in clinical trials, according to the paper. The cost of developing an approvable drug is frequently cited at about $1 billion across 15 years, although recent estimates have ranged as high as $4 billion to $12 billion per drug, depending upon how many of these failures are included in the estimate.
"This basically gives you a computerized safety panel, so someday, when you're deciding among hundreds of thousands of compounds to pursue, you could run a computer program to prioritize for those that may be safest," said Michael Keiser, PhD, co-first author of the paper, who started working on the project as a doctoral student in Shoichet's lab and co-founded SeaChange with Shoichet and John Irwin, PhD, also of UCSF, upon graduation.
It also offers the possibility for identifying possible new uses for medications that are already on the market, according to Peter Preusch, Ph.D., who oversees structure-based drug design grants at the National Institutes of Health's National Institute of General Medical Sciences, which partly supported the study.
"By providing a way to identify the unintended targets of a drug, this advance will not only help streamline the drug development pipeline, but also will provide valuable guidance in efforts to repurpose existing drugs for new diseases and conditions," Preusch said. "This work represents a notable contribution that is likely to find broad applications in the pharmaceutical arena."
The project builds on UCSF's legacy as a leader in developing computer-based approaches to efficiently screen millions of chemicals for those with the best potential for drug development. The UCSF School of Pharmacy was the first to develop computer-based molecular "docking" software, which both public and private researchers use to visualize how potential drugs might attach to target molecules to inhibit their function. It also builds upon UCSF's commitment to industry collaborations that advance pharmaceutical science. Novartis has one of the strongest and most productive drug pipelines in the industry, with more than 130 projects in clinical development, according to the company.
The current project is based on technology UCSF developed, known as the "similarity ensemble approach" (SEA), which compares the shape of each drug to thousands of other compounds and uses that to predict which proteins they might both bind to – essentially, guilt by association. The technique was named among Wired magazine's "Top Scientific Breakthroughs of 2009."
In this project, the UCSF and SeaChange team ran a computer screen on 656 drugs that are currently in clinical use to predict which ones were most likely to bind to the 73 target proteins that appear on Novartis' safety panel for testing drugs for side effects such as heart attacks. Meanwhile, NIBR developed a statistical method of relating those targets to known side effects.
The computer model identified 1,241 possible side-effect targets for the 656 drugs, of which 348 were confirmed by Novartis' proprietary database of drug interactions. Another 151 hits revealed potential side effects that had never been identified for these drugs, yet which Novartis confirmed through lab testing. Among those was a synthetic form of estrogen that has been known for years to cause stomach pain, with no known cause. The screen showed that it binds strongly to a target known as COX-1, which is the protein target of non-steroidal anti-inflammatory drugs, such as aspirin, which also can cause stomach pain, ulceration, and bleeding.
Keiser is co-first author on the Nature paper alongside Eugen Lounkine, PhD, a postdoctoral scholar in the Novartis Institutes for Biomedical Research whose postdoctoral advisors are Urban and Shoichet.
Additional authors include Steven Whitebread, Dmitri Mikhailov and Jeremy Jenkins, from the NIBR's facilities in Cambridge, Mass.; Jacques Hamon, Eckhard Weber and Serge Côté, from NIBR in Basel, Switzerland; and Allison Doak, in the UCSF Department of Pharmaceutical Chemistry.