Novel susceptibility loci identified for osteoarthritis
(HealthDay) -- Five novel single nucleotide polymorphisms (SNPs) are significantly associated with osteoarthritis, including one near the nucleostemin-encoding gene, according to a study published online July 3 in The Lancet.
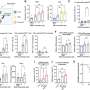
Eleftheria Zeggini, Ph.D., from the Sanger Institute in Cambridge, U.K., and colleagues conducted a large genome-wide association study involving 7,410 unrelated patients with severe osteoarthritis in the arcOGEN study (80 percent of whom had undergone total joint replacement) and 11,009 unrelated controls. The most promising signals were replicated in an independent set of up to 7,473 cases and 42,938 controls. All patients and controls involved in the study were of European descent.
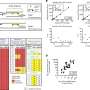
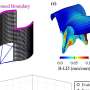
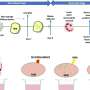
The researchers identified five loci that were significantly associated with osteoarthritis and three loci just below the threshold for significance. The strongest association was seen with rs6976 on chromosome 3 (odds ratio, 1.12), which is in perfect linkage disequilibrium with rs11177, an SNP that encodes a missense polymorphism within the nucleostemin-encoding gene GNL3. In functional studies, patients with osteoarthritis had increased levels of nucleostemin in chondrocytes. Other significant loci were identified on chromosomes 9 and 6, and two loci were identified on chromosome 12. One of the signals close to significance was within the FTO gene. All risk variants exerted small effects and were common.
"We have established novel loci represented by common SNPs that confer a slight risk for osteoarthritis and are associated with the clinically important phenotype of total joint replacement," the authors write. "These results provide a basis for functional studies to identify the underlying causative variants, biological networks, and molecular cause of osteoarthritis."
Several of the authors are employed by deCODE.
More information:
Abstract
Full Text (subscription or payment may be required)
Editorial (subscription or payment may be required)
Copyright © 2012 HealthDay. All rights reserved.