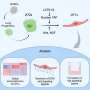
Mathematical modelling to tackle metabolic diseases

Predictive mathematical models of signalling pathways are powerful biological tools that could be used for drug development. Using a similar approach, European scientists developed a computational model for answering research questions regarding the AMP-activated protein kinase pathway.
AMP-activated protein kinase has a master regulatory role in monitoring the cellular energy status. The signalling pathway involving AMP-activated protein kinase controls energy production and consumption, thereby affecting most intracellular biological processes.
The EU 'Systems biology of the AMP-activated protein kinase pathway' (Ampkin) project was designed to contribute to our understanding of how the AMP-activated protein kinase pathways operate. More specifically, project scientists planned to generate predictive kinetic mathematical descriptions of pathway activation/deactivation in order to identify potential drug targets to treat human metabolic diseases.
Using existing data of protein, mRNA expression and metabolite levels enabled scientists to capture the pathway's dynamics and design kinetics models. Comparison of the yeast and mammalian pathways indicated that AMP-activated protein kinase has similar targets and physiological roles in both systems.
Additionally, assay tools were generated for the majority of the steps of the AMP cascade, thereby maximising the use of real data in the mathematical model. Combined with quantitative dynamic datasets generated following activation and deactivation of the AMP-activated protein kinase pathway, it was possible to build mathematical models for the yeast homologue, Snf1.
Importantly, the Ampkin model was designed to assess system perturbations and potentially be used for drug screening. By integrating modelling with experimentation, project partners managed to continuously improve the AMP-activated protein kinase model. This allowed them to address research-related questions and hopefully provide answers for metabolic diseases such as obesity and type II diabetes.