Storm of 'awakened' transposons may cause brain-cell pathologies in ALS, other illnesses
A team of neuroscientists and informatics experts at Cold Spring Harbor Laboratory (CSHL) reports important progress in an effort to understand the relationship between transposons – sequences of DNA that can jump around within the genome, potentially causing great damage – and mechanisms involved in serious neurodegenerative disorders including ALS (amyotrophic lateral sclerosis, also known as Lou Gehrig's disease), FTLD (frontotemporal lobar degeneration) and Alzheimer's disease.
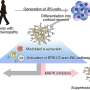
A close analysis of previously unanalyzed genome data has led CSHL Associate Professor Joshua Dubnau, Ph.D., and colleagues to discover a signature of disease that may help explain these and other neural pathologies. As reported by the team September 6 in the journal PloS One, this signature leads them to hypothesize that dormant transposons awaken in the genome, setting off the equivalent of a transposon storm in some brain cells, capable of causing cell death.
Transposons – often called transposable elements, or TEs, by scientists – are understood collectively to occupy a large fraction (~40%) of the genetic material of multicellular organisms, including humans. Most TEs are genomic fossils, effectively inactive. A minority of TEs capable of activation are ordinarily suppressed by a variety of cellular defense mechanisms that have evolved along with life over eons of time to prevent sudden rearrangements (i.e., mutations) of the genetic material.
Dubnau's team, in collaboration with the team of CSHL Assistant Professor Molly Hammell, Ph.D., a bioinformatics expert and co-senior author of the new paper, were intrigued by recent research indicating that some transposable elements are active in brain cells during neurogenesis, the process in which new neurons are born, as well as during normal brain development.
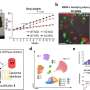
They postulate that mobile chunks of DNA somehow evade the mechanisms that normally restrain them, in certain neuropathologies. For this reason they decided to comb vast datasets for evidence of a particular kind of interaction: the binding of a protein called TDP-43 to RNA "messages" copied from portions of the genome occupied by TEs.
TDP-43 is a versatile protein, known to bind to both DNA and RNA in cells. Accumulations of proteins and RNAs that prominently include TDP-43 are known to have a causal role in ALS pathology. TDP-43 is also known to be associated with FTLD, a condition in which frontal and temporal lobes of the brain atrophy. "Accumulations of TDP-43 in what we call 'inclusion bodies' within the cell's cytoplasm is a shared pathological hallmark in a broad spectrum of neurodegenerative disorders including in ALS and FTLD," Dubnau says.
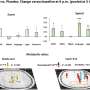
The CSHL scientists noted that TDP-43 was bound to transposon-derived RNAs across three independent, publicly archived genomic datasets from rat, mouse and human samples. Separate research revealed that this association of the binding protein and TE-derived transcripts was less common in the brain cells of FTLD patients, relative to healthy people. They also observed in two different mouse models with TDP-43 dysfunction that the number of observed TE-derived RNA messages was significantly elevated.
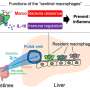
"Putting all these observations together," Dubnau says, "we are now suggesting that TDP-43 normally functions to silence or repress the expression of potentially harmful transposons. When TDP-43 function is compromised, these mobile elements become overexpressed. And this, we hypothesize, may contribute to toxic effects underlying pathologies we see in ALS, FTLD, and perhaps other neurodegenerative conditions."
Current research by the CSHL team aims to flesh out these relationships, and particularly to determine, in Dubnau's words, "whether the unleashing of transposons is a cause or a consequence of the neurodegeneration that we know to be associated with TDP-43 accumulation, in these devastating illnesses."
More information: "Transposable Elements in TDP-43-mediated Neurodegenerative Disorders" appears September 6, 2012 in PloS One. The authors are: W. Li, Y. Lin, L. Prazak, M. Hammell and J. Dubnau. www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0044099