Study reveals rate at which key genetic deletions contribute to male infertility
A large-scale analysis of Y chromosomes from more than 20,000 men finds that two spontaneously recurring deletions along a complex region of the Y chromosome are responsible for approximately 8% of cases of failed sperm production.
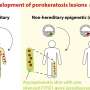
Although previous research had identified deletions in the region of the Y known as AZFc (for azoospermia factor c) as causing severe spermatogenic failure (SSF), this latest analysis, conducted by Whitehead Institute Director David Page and colleagues, is the first to determine how prevalent these deletions are in the general population.
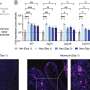
According to the study, published in the November issue of the American Journal of Human Genetics, the deletion known as b2/b4 is found in one of every 2,300 men, increases the risk of SSF 145 times, and is responsible for roughly 6% of cases.
"This deletion almost always results in spermatogenic failure, so it would be extremely rare for it to be transmitted from father to son without medically assisted reproduction," says Page. "Because of this, we can conclude that its prevalence in the population essentially reflects the rate at which this deletion arises spontaneously in men."
"Medically relevant population genetics studies are well established for most of the human genome, but this is the first study of this kind for the Y chromosome," says Steven Rozen, an associate professor at Duke-NUS Graduate Medical School Singapore and first author of the study.
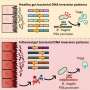
Page notes that this study would not have been possible without the unique sequencing method he developed with collaborators at Washington University in St. Louis to help navigate the structural complexities of the Y chromosome. As Page reported years ago, the Y comprises several regions of large palindromes—areas of mirror-imaged genetic sequences. Such regions render conventional sequencing approaches incapable of detecting extremely subtle genetic differences found hidden among the "mirrors." In response, Page and colleagues developed an approach known as SHIMS (single-haplotype iterative mapping and sequencing) to establish a definitive reference DNA sequence of the Y chromosome.
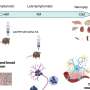
Such structural complexity is not exclusive to the Y chromosome, however. Smaller "halls of mirrors" can be found scattered throughout the human genome, and these areas are likely to be as unstable and prone to mutation as those on the Y chromosome. While the effects of the known deletions of the AZFc region appear to be limited to sperm production, substantially more harmful health effects are apt to arise from mutations elsewhere. Given the inherent challenges of obtaining accurate and complete DNA sequences of mirrored regions, Page believes that the current reference sequence of the human genome is missing potentially meaningful detail—and that the time has come to apply SHIMS broadly.
"The key to SHIMS starts with the realization that there are areas of the human genome that are almost perfectly mirrored repeated sequences that are greater than 99% identical," says Page, who is also an investigator of the Howard Hughes Medical Institute. "When you assemble a sequence from multiple unrelated chromosomes, as was done with the human genome, you cannot make sense of minute but critical differences."
"The human genome reference is a consensus sequence, which is a politically wonderful outcome," he adds. "But in mirrored regions, consensus doesn't really represent anything. A complete and accurate assembly of the human genome will answer questions we do not even know to ask."
More information: American Journal of Human Genetics, November 2, 2012