Text mining: Technology to speed up Alzheimer's biomarker discovery

New research proves that 'text mining' or using the power of computers to read the entire biomedical knowledge base, is a promising new tool in the search for Alzheimer's disease biomarkers.
The research, led by King's College London's Institute of Psychiatry and published in BioMed Central's open access journal, the Journal of Translational Medicine, is the first to use this technology on such a large scale to look for potential biomarkers for disease.
Alzheimer's is the most common form of dementia, and currently affects approximately 500,000 people in the UK. During the course of Alzheimer's disease, 'tangles' or 'plaques' develop in the brain leading to the death of brain cells. However, how and why these develop is still poorly understood and is likely to be due to a combination of factors, including age, genetics and environment. Identifying reliable biomarkers, or biological indicators, for the disease is important for developing early diagnostic tests, and finding new therapies.
Professor Simon Lovestone, lead author of the paper and Professor of Old Age Psychiatry at King's College London's Institute of Psychiatry says: "To our knowledge, this is the first time text mining has been used on this scale in the hunt for biomarkers. Essentially, we used the power of computers to 'read' everything that has ever been written in all biomedical science. We prove that text mining works, and we will take this forward in our hunt for Alzheimer's biomarkers. Our results also demonstrate the value of large data in biomedical science; you could go beyond Alzheimer's disease and use the same approach for other conditions where biomarkers are needed, from cancer to diabetes."
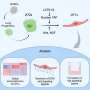
Researchers at King's worked with international colleagues and BioWisdom (now Instem Scientific) to develop a series of 'axioms', or statements, about what a blood biomarker might look like. They then turned this into computer code and by using textual and linguistic analysis, searched for relevant information in all publically available databases, combining neuro-imaging, genetic and proteomic data.
This derived a total of 25 potential biomarkers. The team then validated these - some had previously been identified as potential biomarkers, and in two other cases, they examined the proteins against large sample sets, and showed that the computer approach was correct.
Professor Lovestone, who is also Director of the National Institute for Health Research Biomedical Research Centre (NIHR BRC) for Mental Health at the South London and Maudsley NHS Foundation Trust and King's College London adds: "So far, our search for Alzheimer's disease biomarkers has focused either on an 'omics approach looking at as many proteins or genes as possible, or using a candidate approach looking for the obvious things. However, despite substantial international effort, neither has proved satisfactory. This technology offers an exciting and powerful new tool to advance our research in this field."
Dr Jane Reed, Director of Life Sciences at Instem Scientific and co-author of the paper, says: "This research is a great example of academic-industry collaboration and shows the power of a translational approach to re-use current and legacy data. There is a demand for better methods to predict biomarkers, and this paper validates our in silico approach to biomarker discovery in human disease."
More information: Greco, I. et al. 'Alzheimer's disease biomarkers discovery using in silico literature mining and clinical validation' Journal of Translational Medicine doi:10.1186/1479-5876-10-217