Microevolutionary analysis of Clostridium difficile genomes to investigate transmission
Over recent years, hospital-acquired Clostridium difficile infections have been a significant problem in UK hospitals and globally. There have been concerns that infections may be due to transmission between symptomatic patients, either directly, or indirectly via hospital staff; these concerns were strengthened when enhanced infection control was introduced in England in 2007, and the incidence of C. difficile infection declined. A recent study published in the open access journal Genome Biology, published by BioMed Central, took a genomics approach to assess the incidence of patient-to-patient transmission of C. difficile. The study was supported by the National Institute of Heath Research Oxford Biomedical Research Centre – a collaboration between Oxford University Hospitals NHS Trust and Oxford University.
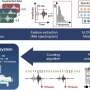
The team, led by Tim Peto and Rosalind Harding at the University of Oxford, sequenced the genomes of C. difficile isolated from 486 patients treated at four hospitals in Oxfordshire between 2006 and 2010. By counting the number of genetic differences between different isolates and estimating the mutation rate of the bacteria, the researchers were able to determine the likely time at which any two isolates became genetically separate and thus, whether the two patients in question could have plausibly caught the infection from each other in the hospital. In other words, genetic divergence implies a time-scale that can be used for judging the likelihood of direct transmission.
The results of the study indicated that, although transmission between patients is likely to occur, it actually happens at relatively low frequency. In particular, concerns that healthcare teams were spreading infection between different hospitals seem to be misplaced. One exception to this general finding is that there were a large number of cases of infection from one particular strain that does appear to have been due to patient-to-patient transmission, emphasising the epidemic nature of this lineage. Notably, this strain has declined in UK hospitals in the last five years.
Dr Xavier Didelot, the study's lead author, said: " This research opens up very exciting opportunities for better understanding how bacterial infections are spread, and what we can do to stop them. The reduced cost of sequencing whole bacterial genomes means we now have the technology for identifying very recent transmissions of infection. Moreover, we can apply this technology even in cases when infection control teams have no suspicion that routes of contact between patients might exist".
More information: Microevolutionary analysis of Clostridium difficile genomes to investigate transmission Xavier Didelot, David Eyre, Madeleine Cule, Camilla Ip, Azim Ansari, Dai Griffiths, Alison Vaughan, Lily O'Connor, Tanya Golubchik, Elizabeth Batty, Paolo Piazza, Daniel Wilson, Rory Bowden, Peter Donnelly, Kate Dingle, Mark Wilcox, Sarah Walker, Derrick Crook, Tim Peto and Rosalind Harding Genome Biology (in press)