Gene decoding obeys road traffic rules
One of life's most basic processes—transcription of the genetic code—resembles road traffic, including traffic jams, accidents and a police force that controls the flow of vehicles. This surprising finding, reported recently by Weizmann Institute researchers in Nature Communications, might facilitate the development of a new generation of drugs for a variety of disorders.
Transcription indeed involves a step resembling the motion of a vehicle: Enzymes "ride" along gene "tracks," creating molecules that will later be translated into the various proteins involved in the life of the cell. In the new study, a research team headed by Prof. Rivka Dikstein of the Biological Chemistry Department has found that just as on the road, maintaining a reasonable distance between the vehicles—that is, the transcribing enzymes—is the surest way to reach a destination safely. In addition to Dikstein, the team included Dr. Nadav Marbach-Bar, Amitai Ben-Noon, Shaked Ashkenazi, Ana Tamarkin-Ben Harush, Dr. Tali Avnit-Sagi and Prof. Michael Walker.
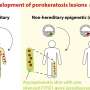
The scientists tracked the transcription of genes coding for tiny regulatory molecules called microRNAs. Working with human cells, they experimented with different rates of transcription: a high rate, in which the enzymes are launched in bursts, and a low one, in which the enzymes are launched individually, at greater intervals. The experiments yielded a paradoxical finding: When the transcription enzymes were launched in bursts, the amount of the resultant microRNA dropped; conversely, when the enzymes were launched at greater intervals, production of microRNA was more efficient.
It turned out that when the enzymes were launched in bursts, one rapidly following the next, they ended up in a traffic jam: When the first enzyme paused at a "road bump"—a molecular signal that creates a pause in transcription—the enzymes that followed crashed into it, falling off the gene. Naturally, such "traffic accidents" reduced the amount of resultant microRNA. In contrast, when the enzymes were launched one by one, they maintained a safe distance: Each had sufficient time to slow down at the "bump" and to succeed at creating a microRNA molecule. In other words, the lower rate of release of individual enzymes proved to be a more efficient method for creating microRNAs.
Because these findings shed new light on the manufacture of microRNAs, they might help in the design of drugs based on these molecules. Discovered as recently as in the 1990s, microRNAs hold great promise for serving as future therapeutics because they can help control gene expression—for example, blocking the activity of cancer-causing genes. This ability is particularly valuable when a molecular process needs to be manipulated at the deepest possible level, inside the cell nucleus.
In a more fundamental sense, the new study helps reveal how transcription is regulated. For example, the study has shown that in inflammation, when the body is threatened with invasion by a virus or bacterium, the release of anti-inflammatory microRNAs is temporarily suspended. The suspension occurs because inflammation increases the launch rate of transcription enzymes, creating traffic jams that reduce the production of the microRNA. This reduction, in turn, "buys time" for the inflammation, giving it a chance to perform its healing function before it is terminated by the microRNA.
Finally, this study helps explain an earlier finding in Dikstein's lab: In longer genes, transcription enzymes tend to be launched at a low rate, that is, at great intervals. The longer the gene, the greater the risk that it has molecular "bumps" that can create traffic jams, derailing transcription. Therefore, transcription enzymes riding along such genes at a lower rate can do their job more efficiently than the enzymes launched in rapid bursts.