Study shows microRNAs can trigger lymphomas
A small group of immune-regulating molecules, when overproduced even moderately, can trigger the blood cancers known as lymphomas, according to a new study led by scientists from The Scripps Research Institute (TSRI).
The six "microRNA" molecules were already known to be overproduced in lymphomas and in many other human cancers, but no one had demonstrated that they can be the prime cause of such cancers—until now. The new study also identified the major biological pathways through which these microRNAs ignite and maintain cancerous growth.
"We were able to show how this microRNA cluster can be the main driver of cancer, and so we now can start to think about therapies to combat its effects," said TSRI Assistant Professor Changchun Xiao. Xiao was the senior investigator for the study, which appeared this week in an advance online version of the EMBO Journal, a publication of the European Molecular Biology Organization.
'Dimmer Switches'
Discovered only in the 1990s, microRNAs are short molecules that work within virtually all animal and plant cells. Typically each one functions as a "dimmer switch" for one or more genes; it binds to the transcripts of those genes and effectively keeps them from being translated into proteins. In this way microRNAs can regulate a wide variety of cellular processes.
The focus of the new study was a cluster of six microRNAs known as miR-17~92, encoded by a single gene on chromosome 13. Studies of miR-17~92, including one from Xiao's lab earlier this year, have shown that it controls various immune-related and developmental processes, depending on the type of cell in which it is expressed.
But the miR-17~92 cluster is best known as a suspected cause of cancers, so much so that it has been dubbed "oncomir-1." Since 2005, scientists have found the cluster to be overproduced in lymphomas, leukemias, brain cancers, breast cancers, prostate cancers and other tumor types. It appears to play an especially prominent role in lymphomas. In a study reported last year, National Cancer Institute researchers found a drastic overexpression of the miR-17~92 cluster in every tumor they sampled from patients with a common type of non-Hodgkin's lymphoma called Burkitt lymphoma.
Researchers have found evidence that this overexpression of miR-17~92 isn't merely an incidental result of cancerous change in cells; it also works to speed up cancerous growth. "What hasn't been known is whether miR-17~92 can be the primary trigger of such cancers," said Xiao.
Identifying a Primary Trigger for Cancer
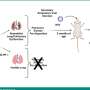
In the new study, he and his colleagues demonstrated that it can be. The project started with a colony of genetically engineered mice that Xiao established several years ago, while doing postdoctoral research in the laboratory of renowned immunologist Klaus Rajewsky at Harvard Medical School. "The mice contain an artificial gene segment that we can activate to overproduce miR-17~92 in any chosen cell type," explained Xiao. In this case, the overproduction occurs only in antibody-producing immune cells called B cells—the same cells from which Burkitt lymphoma originates.
After moving to TSRI to set up his own laboratory in 2008, Xiao expanded this transgenic mouse colony and began to gather data on it. "We found that 80 percent of these mice develop lymphomas within one year," said Hyun-Yong Jin, a graduate student in the Xiao laboratory who was a lead author of the new study.
"It was striking that this very high rate of lymphoma came from only a three-to-fivefold overexpression of miR-17~92 in B cells, whereas human Burkitt lymphomas typically show more than tenfold overexpression," Xiao said.
Having established that miR-17~92 overexpression can powerfully trigger B cell lymphomas, Xiao and his colleagues looked at this microRNA cluster's role in a standard mouse model of Burkitt lymphoma. The B cells of these mice are engineered to overexpress a cancer-inducing "oncogene" called myc, whose hyperactivity—a characteristic of human Burkitt lymphoma cases—triggers a number of abnormalities, including the overproduction of miR-17~92.
The miR-17~92 overproduction turned out to be crucial for the development of these lymphomas. "Deleting miR-17~92 from the B cells of these mice significantly delayed the development of lymphomas and extended the mice's survival," said Maoyi Lai, a research associate in the Xiao laboratory who was a lead author of the study with Hiroyo Oda, a research associate in the Xiao laboratory during the study and now a member of the National Center for Global Health and Medicine in Chiba, Japan. "Looking more closely, we found that the lymphomas that did develop in these mice originated only from B cells in which miR-17~92 had managed to escape deletion and was still being overproduced."
Taking Off the Brakes
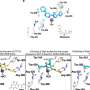
The next step was to investigate how miR-17~92 triggers cancer so powerfully. Using a new technique for finding the binding sites of microRNAs on messenger RNAs, Xiao's collaborator Bryan R. Cullen and colleagues at the Duke University School of Medicine identified hundreds of genes that miR-17~92 works to suppress. A large fraction of these turned out to be genes that normally keep the brakes on cell growth and survival programs. By suppressing these braking genes, miR-17~92 ends up strongly promoting cell growth and survival.
"It affects so many important pathways that even a modest miR-17~92 overexpression apparently moves the cell from a normal growth and survival mode into the cancerous state," Xiao said.
Xiao's team demonstrated the importance of two of these growth/survival pathways by injecting chemical inhibitors of the pathways into mice with miR-17~92-driven lymphomas. "Each inhibitor shrank the tumors and prolonged mouse survival," said Xiao. "We're now studying the effect of combining inhibitors of these miR-17~92-driven cancer pathways and possibly targeting miR-17~92 microRNAs directly."
More information: "MicroRNA-17~92 plays a causative role in lymphomagenesis by coordinating multiple oncogenic pathways," www.nature.com/emboj/journal/v … l/emboj2013178a.html