Recognizing cancer diseases at an early stage: Researchers develop label-free automatic cancer diagnostics
Researchers at the Ruhr-Universität Bochum (RUB) have developed a new spectroscopic method to support pathologists in diagnosing cancer. In the Journal of Biophotonics and the Analyst they compared conventional procedures for colon cancer identification with a novel method called label-free spectral histopathology. "Contrary to previous methods we no longer have to stain the tissue in order to detect cancer," says Professor Klaus Gerwert from the Protein Research Unit Ruhr within Europe (PURE) at the RUB. "In the future, this will give us the opportunity to classify a tissue sample automatically as being either normal or diseased."
Diagnosis: Colon cancer
Today pathologists slice tissue obtained from biopsies into thin sections, stain them chemically, and eventually identify colon cancer by visual inspection under the microscope. This is usually done at an advanced stage of the disease, and the method provides no information about the molecular causes of the tumour. However, the method of spectral histopathology (SHP) established at the RUB Department of Biophysics captures molecular alterations directly in the tissues, especially changes of proteins. It works without any labelling agents, such as fluorescent dyes. SHP may even detect alterations occurring in early tumour stages. Since the analysis uses light beams, SHP is not limited to thin sections of biopsy specimens – in fact, one can apply the method directly in live tissue, where it allows to inspect a site of interest with the aid of fibre-optics. "In the future, we intend to work together with clinical partners and apply spectral histopathology to patients directly via endoscopes," says Klaus Gerwert.
How spectral histopathology works
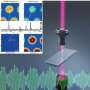
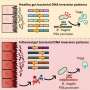
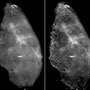
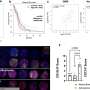
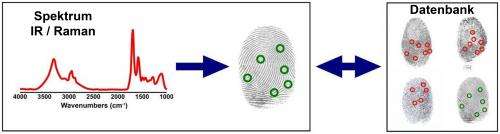
In SHP, researchers record spatially resolved vibration spectra of a tissue using either an infrared or a Raman microscope. A vibration spectrum reflects the condition of all proteins in a tissue at the site measured. Alterations induced by cancer are reflected in the respective spectrum. The spectrum is thus representative of the status of the sample, just like a fingerprint is characteristic of an individual person. Approximately ten million infrared spectra are usually recorded to produce one single tissue image. Using sophisticated computational image analysis procedures, researchers compare these spectra with a reference database. This database contains spectra of already known tissues and tumours, and has been established in the PURE consortium as a collaboration between pathologists, biophysicists and bioinformaticians. The analytical programme allocates each spectrum to tissue types that have been stored in the database, represented by a specific colour—just like an offender who can be identified by comparing his fingerprints with previous database entries. This produces a spatially resolved annotated image of the colon tissue section. Both PURE members, Professor Andrea Tannapfel, Director of the Pathology Institute at the RUB, and Professor Axel Mosig, Head of Bioinformatics at the Department of Biophysics, made the essential contributions in creating the database and the evaluation algorithm. By now, the evaluation programme will run on any commercial laptop.
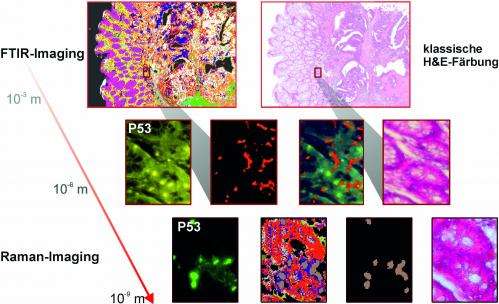
Comparison with conventional tumour detection methods
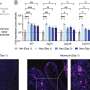
In order to test the sensitivity and specificity of spectral histopathology, the RUB team compared SHP with classical immunohistochemical methods, in which tumours are identified with the aid of fluorescent labels. "The results match perfectly. It demonstrates impressively that spectral histopathology is capable of determining changes in tissue composition with high sensitivity and in an automated fashion," says Prof Gerwert. In fact, sensitivity and specificity of the method exceed 95 per cent. By extending their method to include Raman imaging, the RUB team achieved a higher spatial resolution than they could with infrared imaging, however, at the cost of prolonged measurement time. "Both methods complement each other excellently," comments Klaus Gerwert. "Infrared spectroscopy gives you a rapid overview of the entire tissue section. We can then analyse suspicious regions in more detail by applying Raman imaging." Raman analysis, for example, reveals altered nuclei which are characteristic of tumours.
More information: Kallenbach-Thieltges, A. et al. (2013): Immunohistochemistry, histopathology and infrared spectral histopathology of colon cancer tissue sections, Journal of Biophotonics. DOI: 10.1002/jbio.201200132
Mavarani, L. et al. (2013): Spectral Histopathology of colon cancer tissue sections by Raman imaging with 532 nm excitation provides label free annotation of lymphocytes, erythrocytes and proliferating nuclei of cancer cells, Analyst. DOI: 10.1039/C3AN00370A