Scientists create the first large catalog of interactions between drugs and proteins
The three-dimensional structures of proteins determine how, when and where they bind to drugs and other compounds. In 2012 alone, thousands of structures like this were resolved. Now this mass of information needs to be translated into a biological context that can be used to extract relevant functions from these interactions, as well as significant pharmacological and disease-related effects.
A Spanish National Cancer Research Centre (CNIO) work, led by Alfonso Valencia, Vice-Director of Basic Research, and Michael L. Tress, a researcher on his team, brings together the biggest collection of interactions between pharmacological molecules, including other compounds, and proteins, in the latest edition of the journal Nucleid Acids Research.
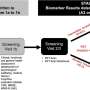
In total, the catalogue includes 16,600 compounds, of which 1,300 contain pharmacological descriptions, and 500,000 interactions that witness the extensive social network that governs the functioning of organisms. The information is available to the entire scientific community via the public FireDB database.
The article's authors, including Paolo Maietta, the first author and a researcher in Valencia's group, point out that: "the creation of FireDB is a reference point for the study of mutations associated with cancer and their biological consequences". According to the researchers, FireDB makes it possible to distinguish the mutations that affect interactions between molecules related to cancer and their effects on the functioning of the cellular machinery.
"For the first time in a single database, this collection of interactions brings together biological information that will be key in understanding the molecular bases of the therapeutic action of drugs, their possible adverse reactions or the creation and development of new therapeutic agents", says Valencia.
More information: FireDB: a compendium of biological and pharmacologically relevant ligands. Paolo Maietta, Gonzalo Lopez, Angel Carro, Benjamin J. Pingilley, Leticia G. Leon, Alfonso Valencia, Michael L. Tress. Nucleic Acids Research (2013). DOI: 10.1093/nar/gkt1127