'Barcode' profiling enables analysis of hundreds of tumor marker proteins at once
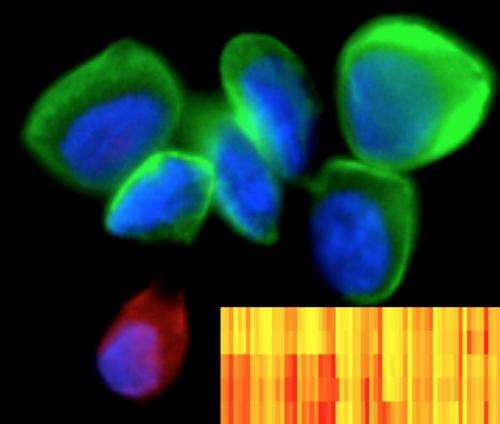
A new technology developed at the Massachusetts General Hospital (MGH) Center for Systems Biology (CSB) allows simultaneous analysis of hundreds of cancer-related protein markers from miniscule patient samples gathered through minimally invasive methods. This new technology uses antibodies linked to unique DNA 'barcodes' to detect a wide range of target proteins and is both powerful and exquisitely sensitive. It could serve as an invaluable tool for helping clinicians gain significant insights into the biology of cancer progression as well as determine why certain cancer therapies stop working or are ineffective to begin with. Development of the technology is reported in the January 15 issue of Science Translational Medicine.
Minimally invasive techniques – such as fine-needle aspiration or circulating tumor cell analysis – are increasingly employed to track treatment response over time in clinical trials, as such tests can be simple and cheap to perform. Fine needle aspirates are also much less invasive than core biopsies or surgical biopsies, since very small needles are used. The challenge has been to comprehensively analyze the very few cells that are obtained via this method. "What this study sought to achieve was to vastly expand the information that we can obtain from just a few cells," explains Cesar Castro, MD, of the MGH Cancer Center and CSB, a co-author of the Science Translational Medicine paper. "Instead of trying to procure more tissue to study, we shrank the analysis process so that it could now be performed on a few cells."
Up until now, pathologists have been able to examine only a handful of protein markers at a time for tumor analyses. But with this new technology, researchers at CSB have demonstrated the ability to look at hundreds of markers simultaneously down to the single-cell level. "We are no longer limited by the scant cell quantities procured through minimally invasive procedures," says Castro. "Rather, the bottleneck will now be our own understanding of the various pathways involved in disease progression and drug target modulation."
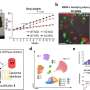
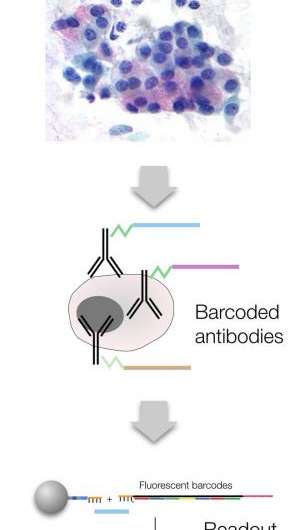
The novel method centers on an approach known as DNA-barcoded antibody sensing, in which unique DNA sequences are attached to antibodies against known cancer marker proteins. The DNA 'barcodes' are linked to antibodies with a special type of glue that breaks apart when exposed to light. When mixed with a tumor sample, the antibodies seek out and bind to their targets; then a light pulse releases the unique DNA barcodes of bound antibodies that are subsequently tagged with fluorescently-labeled complementary barcodes. The tagged barcodes can be detected and quantified via imaging, revealing which markers are present in the sample.
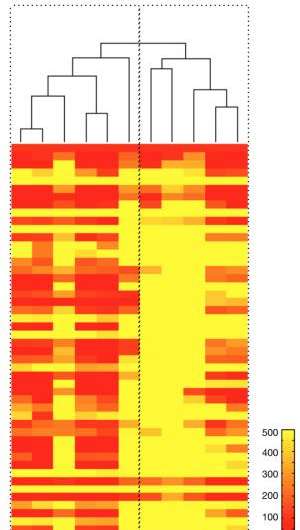
After initially demonstrating and validating the technique's feasibility in cell lines and single cells, the team went on to test it on samples from patients with lung cancer. The technology was able to reflect the great heterogeneity – differences in features such as cell-surface protein expression – of cells within a single tumor and to reveal significant differences in protein expression between tumors that appeared identical under the microscope. Examination of cells taken at various time points from participants in a clinical trial of a targeted therapy drug revealed marker patterns that distinguished those who did and did not respond to treatment.
"We showed that this technology works well beyond the highly regulated laboratory environment, extending into early-phase clinical trials," says Castro, a medical oncologist in the MGH Cancer Center and director of the Cancer Program within the CSB. "Ultimately, the implications for this type of technology could be vast. In this era of personalized medicine, we could leverage such technology not only to monitor but actually to predict treatment response. By obtaining samples from patients before initiating therapy and then exposing them to different chemotherapeutics or targeted therapies, we could select the most appropriate therapy for individual patients."
More information: "Cancer Cell Profiling by Barcoding Allows Multiplexed Protein Analysis in Fine-Needle Aspirates," by A.V. Ullal et al. Science Translational Medicine, 2013.