Scientists unlock evolution of cholera, identify strain responsible for early pandemics

Working with a nearly 200-year-old sample of preserved intestine, researchers at McMaster University and the University of Sydney have traced the bacterium behind a global cholera pandemic that killed millions – a version of the same bug that continues to strike vulnerable populations in the world's poorest regions.
Using sophisticated techniques, the team has mapped the entire genome of the elusive 19th century bacterium. The findings are significant because, until now, researchers had not identified the early strains of cholera, a water-borne pathogen. The discovery significantly improves understanding of the pathogen's origin and creates hope for better treatment and possible prevention.
Researchers have now confirmed the first of two types of cholera, known as classical, was likely responsible for five of the seven devastating outbreaks in the 1800s, all of which most likely originated in waters of the Bay of Bengal.
That strain of cholera had remained a mystery because researchers were unable to examine samples from early victims. The pathogen thrives in the intestines, never reaching teeth or bones, so remnants of its DNA do not exist in skeletal remains. Despite many known cholera burials, access to historical cholera DNA had seemed impossible since it can only be found in soft-tissue remains.
But graduate student Alison Devault and evolutionary geneticists Hendrik Poinar, Brian Golding and Eddie Holmes—working with a team of other scientists—learned that a remarkable collection of tissue specimens was housed at a medical history museum. The Mütter Museum was established by the College of Physicians of Philadelphia in 1858, after the city itself was devastated by cholera earlier in the century.
Researchers carefully sampled a preserved intestine from a male victim of the 1849 pandemic and extracted information from tiny DNA fragments to reconstruct the Vibrio cholera genome.
The results, currently published in The New England Journal of Medicine, could lead to a better understanding of cholera and its modern-day strain known as El Tor, which replaced the classical strain in the 1960s for unknown reasons and is responsible for recent epidemics, including the devastating post-earthquake outbreak in Haiti.
"Understanding the evolution of an infectious disease has tremendous potential for understanding its epidemiology, how it changes over time, and what events play a role in its jump into humans," explains Poinar, associate professor and director of the McMaster Ancient DNA Centre and an investigator with the Michael G. DeGroote Institute of Infectious Disease Research, also at McMaster University.
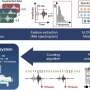
"We need to understand the selective pressures on the pathogen which in turn is driving its evolution, its virulence and hopefully use that information to develop better treatments," he says. Using a sophisticated technique to extract, purify and enrich fragments of the pathogen's DNA, the team collected precious genomic data, which answered many unresolved questions.
The researchers identified "novel genomic islands", or genome regions that don't occur in current strains. In addition, a well-known genic region involved in toxicity of the pathogen (a sequence called "CTX") occurs more times in the ancient strain than in its modern descendants.
This may mean that this strain was more virulent, say researchers, but further testing will be needed.
Regarding the origins, the team's calculations show that the classical strain and El Tor co-existed in humans and estuaries for many centuries, potentially thousands of years prior to the 19th century pandemics, and emerged as a full-blown infection in humans in the early 1800's.
The ancestor of both the classical and El Tor strain likely circulated together in the waters of the Bay of Bengal for several thousand years before emerging in humans during what is known as the first epidemiological transition, or a time of great agricultural revolution and human settlement.
The World Health Organization estimates there are three to five million new cholera cases every year. Of those, 100,000 to 120,000 people typically die from the disease. But with access to historical collections and samples, scientists hope to gain a better understanding of how pandemics arise, spread and ultimately how they might be better controlled or stopped.
"The genomes of ancestral pathogens that have descendants today reside in these archival medical collections all over the world," says Poinar. "We have access to hundreds of thousands of ancient specimens, which hold tremendous potential to determine the origins of past epidemics." Thus these collections represent a treasure trove and should be carefully preserved and maintained.