Scientists discover epigenetic mechanism that could affect risk of obesity-related disease
In one of the largest epigenome-wide association studies (EWAS) to date, published in The Lancet, scientists have identified a new epigenetic mechanism that may play a role in mediating some of the harmful effects of becoming overweight, such as diabetes.
"Obesity increases the risk of heart disease, diabetes, cancer, and a host of other problems, but we know little about the mechanisms by which obesity increases such risk. Genes only explain part of the story", says study leader Professor Nilesh Samani, British Heart Foundation Professor of Cardiology at the University of Leicester, UK.
"Epigenetic changes caused by variation in DNA or environmental factors such as diet, stress, and exposure to chemicals can affect the way genes work (are turned on and off) and may also influence disease susceptibility."
In this study, Samani and colleagues looked at epigenetic changes in DNA in relation to body mass index (BMI), a widely used measure of obesity. One particular type of epigenetic change, a process known as DNA methylation, was examined. DNA methylation involves specific locations along the DNA called cytosine bases, which are modified by the addition of methyl chemical groups.
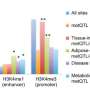
The researchers used microarray technology to measure methylation levels at over 351 000 sites across the genome using whole blood DNA samples taken from 459 individuals of European origin, and identified five sites where the level of methylation correlated with BMI.
These findings were then tested in two additional sets of patients of European ancestry. The results confirmed strong associations with three methylation sites (cg22891070, cg27146050, and cg166772562) located near the HIF3A gene, suggesting that this is a genuine modification of DNA related to changes in weight.
The researchers found that for every 10% increase in methylation at the most significant site—cg22891070—BMI increased by 3.6%, equating to about 0.98 kg/m2 for a person in the original cohort with an average BMI of 27 kg/m2. In comparison, an allele of the known obesity risk gene, FTO, accounts for a more modest 0.39kg/m2 increase in BMI.
They went on to show that changes in methylation at sites in the HIF3A gene also linked with BMI in DNA from fat tissue (a tissue directly involved in obesity) but not from skin DNA, taken from a group of female twins. Further study revealed that changes in methylation of HIF3A were likely to be a result of increased weight rather than a cause.
According to Professor Samani, "The finding of a correlation between HIF3A methylation and BMI was quite unexpected. HIF3A is a component of a protein, hypoxia inducible factor (HIF), which senses oxygen levels in cells and tries to compensate for low levels by affecting the expression of a large number of other genes. To find that the methylation of HIF3A is increasingly altered as someone becomes more obese is remarkable and raises the possibility that HIF may also be involved in mediating some of the deleterious effects of becoming overweight."
He concludes, "Further studies are needed to understand how and when obesity affects methylation at HIF3A and what the consequences are, but the findings could eventually lead to new treatments that may tackle the adverse effects of obesity on health. At a more general level, our study shows that investigating epigenetic changes in DNA may reveal new mechanisms involved in common diseases."
Writing in a linked Comment, Therese Murphy and Jonathan Mill from the University of Exeter, Devon, UK, say, "[This] study represents an important advance for both obesity-related research and the specialty of epigenetic epidemiology. BMI is a good phenotype for population-based epigenomic studies: it is an accurate measure that is routinely collected in most cohort studies. The widespread uptake of [new tools for epigenetic profiling]...means that large collaborative EWAS meta-analyses can be done, building on the success of similar approaches in genetics. Whether EWAS will be as successful for other clinical phenotypes—especially those manifest in more inaccessible tissues such as brain, or more directly affected by confounding factors such as cellular heterogeneity, environmental exposures, and drugs—remains to be seen."
More information: Study paper: www.thelancet.com/journals/lan … (13)62674-4/abstract