Scientists make breakthroughs in ovarian cancer research
Scientists at A*STAR's Institute of Medical Biology (IMB) and the Bioinformatics Institute (BII) have found new clues to early detection and personalised treatment of ovarian cancer, currently one of the most difficult cancers to diagnose early due to the lack of symptoms that are unique to the illness.
There are three predominant cancers that affect women – breast, ovarian and womb cancer. Of the three, ovarian cancer is of the greatest concern as it is usually diagnosed only at an advanced stage due to the absence of clear early warning symptoms. Successful treatment is difficult at this late stage, resulting in high mortality rates. Ovarian cancer has increased in prevalence in Singapore as well as other developed countries recently. It is now the fifth most common cancer in Singapore amongst women, with about 280 cases diagnosed annually and 90 deaths per year .
Identifying Ovarian Cancer Earlier
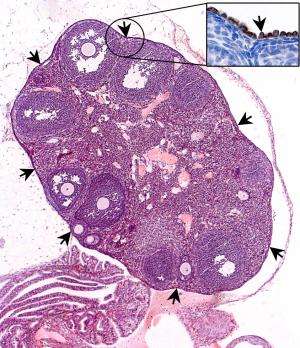
IMB scientists have successfully identified a biomarker of ovarian stem cells, which may allow for earlier detection of ovarian cancer and thus allow treatment at an early stage of the illness.
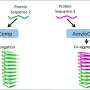
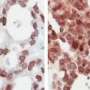
The team has identified a molecule, known as Lgr5, on a subset of cells in the ovarian surface epithelium . Lgr5 has been previously used to identify stem cells in other tissues including the intestine and stomach, but this is the first time that scientists have successfully located this important biomarker in the ovary. In doing so, they have unearthed a new population of epithelial stem cells in the ovary which produce Lgr5 and control the development of the ovary. Using Lgr5 as a biomarker of ovarian stem cells, ovarian cancer can potentially be detected earlier, allowing for more effective treatment at an early stage of the illness (see Annex A). These findings were published online in Nature Cell Biology in July 2014.
Bioinformatics Analysis to Develop Personalised Treatment
Of the different types of ovarian cancers detected, high-grade serous ovarian carcinoma (HG-SOC) is the most prevalent of epithelial ovarian cancers . It has also proven to be one of the most lethal ovarian cancers, with only 30 per cent of such patients surviving more than five years after diagnosis . HG-SOC remains poorly understood, with a lack of biomarkers identified for clinical use, from diagnosis to prognosis of patient survival rates.
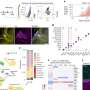
By applying bioinformatics analysis on big cancer genomics data , BII scientists were able to identify genes whose mutation status could be used for prognosis and development of personalized treatment for HG-SOC.
The gene, Checkpoint Kinase 2 (CHEK2), has been identified as an effective prognostic marker of patient survival. HG-SOC patients with mutations in this gene succumbed to the disease within five years of diagnosis, possibly because CHEK2 mutations were associated with poor response to existing cancer therapies (see Annex B). These findings were published in Cell Cycle in July 2014.
Mortality after diagnosis currently remains high, as patients receive similar treatment options of chemotherapy and radiotherapy despite the diverse nature of tumour cells within tumours and across different tumour samples. With these findings, personalised medicine for ovarian cancer could be developed, with targeted treatment that would be optimised for subgroups of patients.
Prof Sir David Lane, Chief Scientist, A*STAR, said, "These findings show how the various research institutes at A*STAR offer their expertise in developing new approaches to examine different aspects of the same disease that have not been successfully studied before, such as ovarian cancer. The diverse capabilities and knowledge of our scientists allows us to investigate diseases holistically, from diagnosis to treatment."
More information: "Lgr5 marks stem/progenitor cells in ovary and tubal epithelia" by Annie Ng et al. Nature Cell Biology, 2014.
"Identification of two poorly prognosed ovarian carcinoma subtypes associated with CHEK2 germline mutation and non-CHEK2 somatic mutation gene signatures" by Ghim Siong Ow, Cell Cycle, www.landesbioscience.com/journ … ls/cc/article/29271/