Overlooked for 30 years: Novel peptide plays role in Alzheimer's disease

A team led by Christian Haass has identified a novel peptide that plays a role in Alzheimer's disease: The previously overlooked eta-amyloid interferes with neuronal function and may antogonize beta-amyloid – a finding that has implications for ongoing clinical trials.
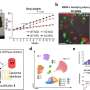
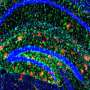
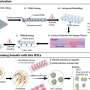
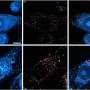
Alzheimer's disease is associated with the appearance of characteristic neurotoxic protein aggregates in various regions in the brain. Chemical analysis of these insoluble deposits reveals that they are made up of a family of short protein fragments, referred to as beta-amyloid peptides, which are derived from a precursor protein called APP by the sequential action of two enzymes. An international team of researchers led by Christian Haass (Professor of Metabolic Biochemistry at LMU and Speaker for the German Center for Neurodegenerative Diseases in Munich) and Dr. Michael Willem (LMU) has now made a discovery which extends this picture of the pathogenesis of Alzheimer's disease, and has potentially far-reaching implications for our understanding of the condition: "A second mode of APP cleavage exists, which involves a previously unknown cleavage and generates an alternative peptide," says Christian Haass.
Its discoverers refer to the newly characterized protein fragment by the Greek letter eta, christening it 'amyloid-η'. "The processing pathway that produces it has been overlooked for 30 years. This is because investigators including myself have focused their attention on elucidating the origins of the beta-amyloid and on attempts to cure Alzheimer's by inhibiting production of this peptide," Haass explains. The paper that describes the generation of amyloid-η appears in the new issue of the journal Nature.
A previously unknown interaction
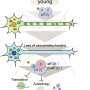
In collaboration with neurobiologist Dr. Hélène Marie based at the IPMC-CNRS in Valbonne (France) and with the local colleagues from the Technical University of Munich (TUM) in the Synergy Excellence Cluster (Professor Arthur Konnerth and Dr. Marc Aurel Busche), the LMU researchers have also studied the effects of the eta-amyloid on nerve-cell function in the brain. Beta-amyloid is known to make nerve cells hyperactive, and now it turns out that the eta-amyloid antagonizes this effect. "So here we have two small peptides snipped from the same precursor protein, which have opposite effects on neuronal activity, and whose actions must normally be carefully balanced," Haass explains.
These findings have immediate implications for ongoing clinical trials in humans, all of which are targeted to beta-amyloid. One of the trials, for instance, is designed to ascertain whether or not pharmacological inhibition of beta-secretase, the proteolytic enzyme that initiates the release of the toxic beta-amyloid from APP, can reduce memory loss in patients with Alzheimer's. Haass, Willem and colleagues confirmed that blocking the action of the beta-secretase does indeed reduce levels of beta-amyloid. However, this is accompanied by a massive increase in the amount of eta-amyloid generated. "This could result in attenuation of neuronal activity and might therefore compromise brain function," says Haass. He therefore suggests that investigators need to be on the look-out for any signs of unanticipated side-effects in the current clinical trials.
More information: η-Secretase processing of APP inhibits neuronal activity in the hippocampus, Nature (2015) DOI: 10.1038/nature14864