Breakthrough in cell-type analysis offers new ways to study development and disease
Like skilled assassins, many diseases seem to know exactly what types of cells to attack. While decimating one cadre of cells, diseases will inexplicably spare a seemingly identical group of neighbors. What makes cells vulnerable or not depends largely on the kinds and amounts of proteins they produce - their "translational profile," in the lingo of molecular biology. For this reason, scientists have struggled to parse the subtle molecular differences among the hundreds of specialized cell types that are tangled together in tissues like the brain.
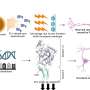
Now, in back-to-back papers in the November 14 issue of the journal Cell, researchers at The Rockefeller University report a breakthrough in cellular analysis that slashes through this Gordian knot. The scientists have developed a method to reveal translational profiles by isolating the genetic messages that govern protein production in different cell types. The new method, translating ribosome affinity purification (TRAP), uses genetically engineered mice to capture these messages as they pass through the protein production factories called ribosomes. Because the mice have been made to express a specially tagged ribosome in only one particular cell type, the TRAP method can identify all the genetic messages that give that cell type its unique identity, including, perhaps, its susceptibility to disease.
So TRAP solves a problem that has been a fundamental barrier to a deeper understanding of the brain and how neurological diseases attack it. But because the method can be used to distinguish any type of cell in any tissue in any organ -- not just brain cells -- it has applications for research into afflictions as varied as cancer metastases, coronary artery disease and diabetes. The work is a collaboration between the labs of Rockefeller professors Nathaniel Heintz and Paul Greengard as well as colleagues at Northwestern University and the Translational Genomics Research Institute (TGen).
"We've created a novel, generally applicable tool that can be used by a broad spectrum of the scientific community," says Heintz, who is the James and Marilyn Simons Professor, head of the Laboratory of Molecular Biology and a Howard Hughes Medical Institute investigator. "I think it will rapidly spread into many of areas of biology."
Greengard, Vincent Astor Professor and head of the Laboratory of Molecular and Cellular Neuroscience, says about half of the research in his lab now employs the new technique to study the biochemical basis of Parkinson's, Alzheimer's and Huntington's diseases, as well as the still-mysterious ways in which psychoactive drugs fight schizophrenia and depression. TRAP should fundamentally change biochemical studies of the brain and the speed at which they yield results, he says.
"We can look at a thousand genes instead of one at a time, so things should clear a thousand times faster," says Greengard, who won the Nobel Prize in Physiology or Medicine in 2000 for research into how neurons communicate.
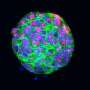
The TRAP method grew out of a project known as GENSAT (for Genetic Expression Nervous System Atlas) that Heintz and Rockefeller professor Mary Beth Hatten launched in 2000 to visualize the contributions of individual genes to the mouse brain. Heintz and his colleagues had developed a technique to engineer large pieces of DNA carried in bacterial artificial chromosomes (BACs), which can insinuate themselves into the genomes of other organisms. They were able to insert the genetic code for green fluorescent protein (EGFP) within the regulatory domain of any gene of interest. When one of these modified BACs is transferred into mice, expression of the EGFP mimics that of the gene of interest, lighting up cells with a green glow that shows researchers all of the cells in which that particular gene functions.
The GENSAT database laid out in glowing green myriad cell types of the mouse brain. And it provided genetic markers for each kind. But it was an accomplishment that was also a taunt. Ultimately, the researchers wanted to go deeper to understand the precise biochemical characteristics of the cell types they had brought into focus, to learn what makes cells vulnerable to attack - and possibly how to protect them from it - by discovering what's unique to the susceptible cells and the ones that are resistant. Enter TRAP.
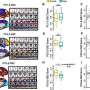
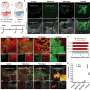
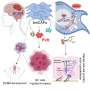
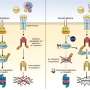
Heintz, postdoctoral fellow Myriam Heiman and colleagues attached an EGFP to the surface of the ribosome and used it as a handle to pick out the cell's protein factories and the genetic messages passing through them, called messenger RNAs (mRNAs). Using the GENSAT techniques and findings, they designed new mouse lines that made tagged ribosomes in each of four different cell types. Heiman and colleagues focused on the brain cells that respond to dopamine, an important neurotransmitter involved in muscle movement and emotion regulation, among other things. They used the handle they had made to pluck out the ribosomes and mRNAs from these brain cells and freeze them within minutes of dissection, preserving the "messages" largely as they were inside the living animal and minimizing degradation. Alternative approaches to getting the profiles of cell types in complex tissues have been disappointing because they require the physical isolation of whole cells from the tissues in which they are embedded. TRAP bypasses that logistical nightmare by going straight for the ribosome.
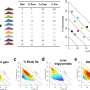
The method proved so sensitive that researchers were able to identify a few hundred genetic messages that differ between two types of dopamine-sensing brain cells that previously had seemed nearly identical. Because these cells are crucial elements of the neural circuit that degenerates in Parkinson's and Huntington's diseases, the newly identified proteins could aid in the design of drugs that would allow these two key cell types to be treated independently of one another.
"We can probe into each cell type, see what is there and possibly identify better therapeutic targets," says Heiman. "This approach is much more in line with a rational drug design."
In a major application of TRAP published as a separate study in the same issue of Cell, Rockefeller scientists led by research associate Joseph Doyle and postdoctoral associate Joseph Dougherty went on to characterize the protein profiles of 24 types of cells in the central nervous system, identifying thousands of proteins that were previously unassociated with known cell types. The work in Cell provides the research community with 16 lines of transgenic mice that can be used for a sweeping range of potential neurological experiments. Heintz says his lab will make many more as they pursue detailed studies of other cell types.
"We can now study the molecular phenotypes that occur in specific cell types in response to genetic, environmental or pharmacological perturbations, determine the precise changes within specific cell types as they progress through development and examine the detailed properties of cells as they succumb to the pathological events occurring in neurological diseases such as ataxia telangiectasia, autism and Rett syndrome," says Heintz. "We are very excited by the opportunities this offers to us and our colleagues for investigation of these issues."
Source: Rockefeller University