SUMO-snipping protein plays crucial role in T and B cell development
When SUMO grips STAT5, a protein that activates genes, it blocks the healthy embryonic development of immune B cells and T cells unless its nemesis breaks the hold, a research team led by scientists at The University of Texas MD Anderson Cancer Center reports today in Molecular Cell.
"This research extends the activity of SUMO and the Sentrin/SUMO-specific protease 1 (SENP1) to the field of immunology, in particular the early lymphoid development of T and B cells," said the study's senior author, Edward T. H. Yeh, M.D., professor and chair of MD Anderson's Department of Cardiology.
SUMO proteins, also known as the small ubiquitin-like modifiers or Sentrin, attach to other proteins in cells to modify their function or to move them within a cell. SENP1 is one of a family of six proteins that snips SUMO off of SUMO-modified proteins. SUMOylation (SUMO modification) of proteins has been implicated in development of cancer, heart and neurodegenerative diseases, among others.
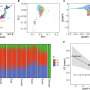
The team first analyzed the role of SENP1 in the development of lymphoids in mice and found it is heavily expressed in precursor cells, the early stages of B and T cell development.
Working with genetically modified mice they developed that lack SENP1 gene expression, Yeh and colleagues found the mouse embryos had severe defects in their T and B cells, white blood cell lymphocytes that identify and fight infection.
SUMO pins STAT5 in the nucleus
Subsequent experiments led them to STAT5, a transcription factor known to play critical roles in the development and function of immune cells. Transcription factors work in the cell nucleus, activating gene expression by connecting to a gene's promoter region.
"STAT5 works in a cycle, moving from the cytosol of a cell into the nucleus to activate genes and then back out to the cytosol," Yeh said. "We found that when STAT5 is SUMOylated in the nucleus it gets trapped there when there's no SENP1 to remove SUMO."
The team found that SUMO muscles in on two other signaling events that govern STAT5 activity - phosphorylation and acetylation.
SUMO inhibits STAT5 signaling
STAT5 is activated in the cell cytosol when the JAK tyrosine kinase attaches a phosphate group at a specific site on the STAT5 protein. This transformed STAT5 crosses the nuclear membrane into the nucleus to transcribe genes.
The team found that SUMO attaches to STAT5 close to its phosphorylation site and that cells lacking SENP1 have increased SUMOylation and decreased phosphorylation.
SUMOylation vs. acetylation
In addition to phosphorylation, acetylation of STAT5 has been shown to be essential for STAT5 to cross the nuclear membrane into the nucleus to enhance gene transcription. Yeh and colleagues found that SUMO competes directly with acetyl groups for the same binding site, inhibiting acetylation.
"Without SENP1 to remove SUMO, STAT5 can't be acetylated or phosphorylated and can't be recycled for use again," Yeh said. "We discovered that SENP1 controls lymphoid development through regulation of SUMOylation of STAT5."
Since Yeh's lab discovered SUMOylation in 1996, SUMO has been found to alter the function of thousands of proteins.
Yeh is hosting the 6th International Conference SUMO, Ubiquitin, UBL Proteins: Implications for Human Diseases Feb. 8-11 in the Dan L. Duncan Building at MD Anderson. Yeh organizes the meeting every other year.
"There used to be so little known about SUMO. Now, a protein is assumed to be SUMOylated until proved otherwise," Yeh said.