Energy levels link sleep control mechanisms

Sleep, or lack of it, can determine level of cognitive performance which is linked with accidents as well as increased risk of serious health problems. Links between cell energy levels, gene transcription and sleep rhythms may uncover answers to sleep disorders and the ill-effects of sleep deprivation.
Timing and quality of sleep is determined by a homeostatic process that compensates for sleeplessness and a circadian cycle to determine the time of day when we should sleep. The effects of even minor misalignments between these two processes, as in jet lag for example, indicate that sleep control is a very relevant feature in cognitive performance and life quality.
Previous studies by the EU-funded 'Redox potential as an interface between sleep homeostasis and circadian rhythms' (Redoxsleepcircadian) project members have suggested that although the two cycles are supposed to operate independently, the two main clock genes work together for homeostatic sleep regulation and generation of circadian rhythms. Not only that, but both CLOCK and NPAS2, the core clock genes, depend on the redox potential in cells suggesting that the two sleep mechanisms are linked through cell metabolism.
The aim of the recently completed project Redoxsleepcircadian was to understand the cell processes that determine our daytime performance and sleep quality. By inducing circadian rhythms in a type of connective tissue cell, fibroblasts, the sleep scientists could determine if there were any accompanying redox changes.
Using redox genetic probes along with a time lapse imaging system, changes in redox potential were recorded in living fibroblasts. Any increases in transcriptional activity in the clock genes Per1 and Per2 during sleep deprivation may indicate energy deficits in prolonged wakefulness. Per 1 and Per 2 are under direct control of CLOCK and NPAS2.
Results indicated that Per2 regulates activity-induced and circadian mechanisms central to regulation of sleep and wake periods. Moreover, molecular feedback underlying circadian rhythm generation was able to regulate the requirements of homeostatic sleep need.
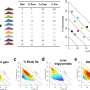
In parallel, the project team also focused on Per2 in mice. This sleep gene has been observed to increase expression after sleep deprivation (SD) in the rodent. The scientists demonstrated that all living mice increased levels of Per2 protein production with different dynamics in the brain. This was particularly so in the cerebral cortex as well as in the liver and the kidney.
Interestingly, extended times without sleep affected Per2 RNA expression through circadian and non-circadian mechanisms always resulting in increased Per2 protein. This suggests that there is no separation between the two cycles.
At the recent close of the project, Redoxsleepcircadian was further investigating the role of the main circadian pacemaker, the suprachiasmatic nucleus (SCN), a miniscule region on the brain's midline. Regulating many neuronal and hormonal activities, the SCN is key player in the relationship between clock genes, sleep and wakefulness.
Ability to control sleep rhythms could mean a welcome reprieve for patients with sleep disorders. Jet lag and disorientation due to shift work for many of the world's labour force could also become things of the past.