New method provides fast, accurate, low cost analysis of BRCA gene mutations in breast cancer
Individuals with mutations in BRCA1 and BRCA2 genes have a significantly higher risk of developing breast and ovarian cancers. Families at risk have been seeking genetic testing and counseling based on their mutation carrier status, but the standard method of direct sequencing is labor-intensive, costly, and it only targets a part of the BRCA1 and BRCA2 genes. A group of Canadian scientists has developed a new sequencing approach to provide a more effective method of BRCA1/2 mutational analysis. Their work is published in the September issue of The Journal of Molecular Diagnostics.
"A comprehensive understanding of BRCA1/2 genotypes and the associated tumor phenotypes is needed to establish targeted therapies," notes lead investigator Hilmi Ozcelik, PhD, of the Fred A. Litwin Centre for Cancer Genetics, Samuel Lunenfeld Research Institute, Mount Sinai Hospital, Toronto, Ontario, Canada. "Recent studies have suggested that certain chemical inhibitors are effective for the treatment of breast cancer in patients with BRCA1/2 mutations. Therefore, availability of new, affordable, and comprehensive technologies to screen for these mutations will be critical to identify patient-candidates for targeted therapies."
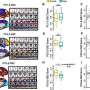
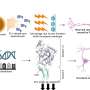
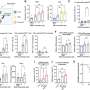
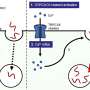
The investigators used a technique called long range PCR to generate amplified BRCA1/2 fragments, known as amplicons, from the DNA of 12 familial breast cancer patients. The amplicons were screened using deep sequencing, also known as Next Generation Sequencing (NGS), which allows for the simultaneous screening of millions of DNA molecules, thereby dramatically increasing speed and throughput. While conventional screening methods target only the exons of BRCA1/2, deep sequencing can screen the entire genomic region, including introns and untranslated regions. The specimens had been previously analyzed using conventional methods, allowing for a comparison of results.
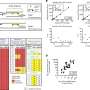
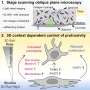
In addition to identifying one genetic variant that was missed due to human error, the new method successfully identified all of the expected BRCA1/2 variants. They identified both exonic and exon/intron boundary variants. The test was done at a very low cost, and with a turnaround time of 12 days. "One of the key advantages of workflow of long-range PCR is the ability to visually detect large genomic duplications, deletions, and insertions," notes Dr. Ozcelik. "When combined with next generation sequencing, long range PCR can be a powerful tool in the detection of BRCA variants in the clinical setting. Our method confirmed the presence of variants with very high accuracy, and without false-positive results."
Long-range PCR and next generation sequencing identified a wide range of intronic BRCA1/2 variants, both commonly occurring and rare, that individually or in combination may impact BRCA1/2 function. Dr. Ozcelik notes that despite a small sample size, the data shows great variability in the number, type, and frequency of variants that can be identified from familial breast cancer patients.
"Our challenge now is to establish analytical methods that systematically investigate this more comprehensive data in order to provide better risk information for clinical management of the disease," says Dr. Ozcelik. "Given the extensive level of genetic information acquired from each patient, profiles can be constructed in breast cancer patients compared to population controls to produce a more effective means of generating BRCA1/2-associated risk to the individuals and their families.
More information: “Long-Range PCR and Next Generation Sequencing of BRCA1 and BRCA2 in Breast Cancer, H. Ozcelik, X. Shi, M.C. Chang, E. Tram, M. Vlasschaert, N. DiNicola, A. Kiselova, D. Yee, A. Goldman, M. Dowar, B. Sukhur, R. Kandel, and K. Siminovitch. DOI: dx.doi.org/10.1016/j.jmoldx.2012.03.006. The Journal of Molecular Diagnostics, Volume 14, Issue 5 (September 2012)