Influenza research: Can dynamic mapping reveal clues about seasonality?
Influenza outbreaks in the United States typically begin with the arrival of cold weather and then spread in seasonal waves across geographic zones. But the question of why epidemics can vary from one season to the next has baffled scientists.
In a paper titled "Deviations in Influenza Seasonality: Odd Coincidence or Obscure Consequence," Elena Naumova, Ph.D., professor of civil and environmental engineering at Tufts School of Engineering, and collaborators from the U.S. and India suggest that the search for answers has been thwarted, in part, by the lack of standardized research methods.
This paper builds on Naumova's previous work in which she suggests a role for dynamic mapping in epidemiological research. Here, the team concludes that newly emerging technologies like dynamic mapping can be used in concert with traditional approaches, which Naumova describes as "fraught with problems." The paper was published in advance of print in Clinical Microbiology and Infection.
Naumova points out several of these problems. Data collection methods are not uniform. Researchers use ambiguous terminology and definitions. Research results are not presented in a uniform manner. "This produces volumes of information—or noise—that is prone to substantial measurement error and uncertainty that potentially obscures the causes behind seasonality rather than illuminating them," she says.
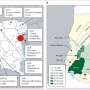
Another way influenza research falls short is that it doesn't take into account the behavior of the disease as it changes across time and geography. Using dynamic mapping as a tool, Naumova, the paper's senior author, and her team analyzed hospitalization records for adults age 65 and over from across the United States during individual flu seasons in 1991, 1997, 1999, and 2003.
The data, provided by the Centers for Medicare and Medicaid Services, were superimposed onto national maps showing average weekly temperatures. The data were then transformed into an interactive movie from which researchers could view data, including origin and intensity of the outbreak, as they appeared over time. Each season exhibited variations in disease patterns.
For example, the 1991 outbreak originated in the Alabama-Louisiana region and then spread north. By contrast, the 1997 the epidemic started in central Tennessee. The 1999 epidemic started in western Minnesota while the 2003 epidemic spread from the gulf region of Texas and Louisiana, according to Naumova. Three of the epidemics peaked in the first week of January while a fourth peaked in the last week of December. Nationally, the average peak occurred during the third week of January.
Naumova points out that the four seasons reveal the diversity exhibited by seasonal outbreaks. "In these years we see that the outbreaks points of origin shifted, as did its geographic spread," she says.
Naumova's ongoing research involves developing mathematical models for analysis of large databases to study infectious diseases and exposure assessment. In an earlier study, researchers analyzed a larger data set from 1991 to 2004. They found that over the 13 seasons, seasonal flus generally peaked earliest in the west and spread east.
While mapping reveals variations in peak timing, intensity, and geography, the factors that influence these parameters are unknown and require further investigation. In the larger picture, Naumova says dynamic mapping could potentially help fill a gap in influenza research.
"It would help give us insight into what kinds of questions to ask. This can lead to a framework from which we are able to develop new research questions," says Naumova, who adds that mapping is only one part of the puzzle. "We also must establish methodologies that use science-based definitions, reliable data and standard methods for presenting data and assessing statistics."
More information: onlinelibrary.wiley.com/doi/10 … 012.03959.x/abstract