New invasive non-Typhoidal Salmonella epidemic identified in sub-Saharan Africa
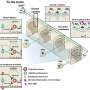
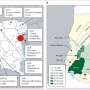
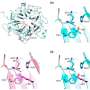
![Geospatial transmission of invasive Salmonella Typhimurium isolates in sub-Saharan Africa. (a,b) Phylogeographic diffusion of lineages I (a) and II (b) across sub-Saharan Africa over time based on a discrete geospatial model with associated geographic coordinates. [DOI: 10.1038/ng.2423] New invasive non-Typhoidal Salmonella epidemic identified in sub-Saharan Africa](https://scx1.b-cdn.net/csz/news/800a/2012/newinvasiven.jpg)
(Medical Xpress)—A new study out today reveals that the emergence and spread of a rapidly evolving invasive intestinal disease, that has a significant mortality rate (up to 45%) in infected people in sub-Saharan Africa, seems to have been potentiated by the HIV epidemic in Africa.
The team found that invasive non-Typhoidal Salmonella (iNTS) disease is caused by a new form of the bacteria Salmonella Typhimurium that has spread from two different focal hubs in Southern and Central Africa beginning 52 and 35 years ago, respectively. They also found that one of the major contributing factors for the successful spread of iNTS was the acquisition of genes that afford resistance to several front line drugs used to treat blood-borne infection such as iNTS.
iNTS is a blood-borne infection that kills approximately one of four people in sub-Saharan Africa who catch it. Yet, in the rest of the world, NTS is a leading cause of acute inflammatory diarrhoea that is self-limiting and tends to be fatal in less than 1 per cent of people infected. The disease is more severe in sub-Saharan Africa than the rest of the world because of factors such as malnutrition, co-infection with malaria or HIV and potentially the novel genotype of the Salmonella bacteria.
"The immune system susceptibility provided by HIV, malaria and malnutrition at a young age, may provide a population in sub-Saharan Africa that is large enough for this detrimental pathogen to enter, adapt, circulate and thrive," says Chinyere Okoro, joint first author from the Wellcome Trust Sanger Institute. "We used whole genome sequencing to define a novel lineage of Salmonella Typhimurium that is causing a previously unrecognised epidemic across the region. Its genetic makeup is evolving into a more typhoid like bacteria, able to efficiently spread around the human body"
From sequenced samples, the team created a phylogenetic or 'family tree', depicting the pathogen's evolution, dating when each sample first emerged and overlaying this with geographical information about where these samples came from. They found that this invasive disease comprises of two very closely related waves; the first wave originated from a possible south-eastern hub, about 52 years ago and the second originated about 35 years ago, possibly from the Congo Basin.
"The HIV epidemic in sub-Saharan Africa is thought to have begun in a central region and underwent expansion eastwards, a strikingly similar dynamic to that observed for second iNTS wave," says Dr Robert Kingsley, joint first author from the Wellcome Trust Sanger Institute. "Our findings suggest the current epidemic of iNTS and its transmission across sub-Saharan Africa may have been potentiated by an increase in the critical population of susceptible, immune-compromised people."
The team identified that the vast majority of samples from the second wave of iNTS contains a gene that makes them resistant to chloramphenicol, a frontline antibiotic in the treatment of Salmonella. This gene was not present in the samples from the first wave of iNTS. This observation suggests that iNTS acquired this gene early on in the evolution of the second wave, probably around the time of its spread from the Congo basin.
"Because it acquired resistance to chloramphenicol, this pathogen has much greater opportunity to survive and spread across the region," says Professor Gordon Dougan, lead author from the Wellcome Trust Sanger Institute. "This is the first time that the power of whole-genome sequencing has been used to track the spread of iNTS. Our research highlights the power this approach has to monitor the emergence and spread of dangerous pathogens both locally and globally over time."
"There has been some evidence that this disease can be passed from human to human. Now the race is on to discover how NTS is actually transmitted in sub-Saharan Africa so that effective intervention strategies can be implemented."
More information: Chinyere K. Okoro, Robert A. Kingsley, Thomas R. Connor et al. 'Intra-continental spread of human invasive Salmonella Typhimurium pathovariants in sub-Saharan Africa'. Published in Nature Genetics 30 September 2012. DOI: 10.1038/ng.2423