New mechanism of action for PARP inhibitors discovered
New understanding of how drugs called PARP inhibitors, which have already shown promise for the treatment of women with familial breast and ovarian cancers linked to BRCA mutations, exert their anticancer effects has led to the identification of ways in which the patient population that might benefit from PARP inhibitors could be expanded.
Yves Pommier, M.D., Ph.D., chief of the Laboratory of Molecular Pharmacology at the National Cancer Institute's Center for Cancer Research in Bethesda, Md., and colleagues reported these data in Cancer Research, a journal of the American Association for Cancer Research.
"In recent years, drugs classified as poly (ADP-ribose) polymerase (PARP) inhibitors have been shown to be promising anticancer agents for breast and ovarian cancers deficient in either BRCA1 or BRCA2," said Pommier. "Prior to our study, PARP inhibitors were thought to work primarily by blocking the DNA repair function of members of the PARP family of proteins, leading ultimately to cancer cell death."
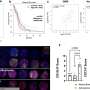
In their initial studies, Pommier and his colleagues found that the PARP inhibitor olaparib was more toxic to cultured cells than genetic elimination of PARP1.
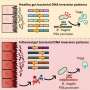
According to Pommier, these results indicated that olaparib must have additional modes of action, and their detailed cellular analyses identified a critical one: olaparib was trapping PARP proteins, specifically PARP1 and PARP2, at sites of DNA damage, and the trapped PARP protein-DNA complexes were highly toxic to cells.
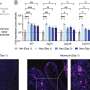
When the trapping ability of olaparib was compared with that of two other PARP inhibitors under clinical development, it was found that the trapping potency of the three drugs differed markedly: niraparib was more potent than olaparib, which was in turn substantially more potent than veliparib. In contrast, olaparib was the most potent inhibitor of DNA repair function, followed by veliparib and then niraparib.
"Critical to this study, is the demonstration that PARP inhibitors are not equivalent with respect to their potency to trap PARP proteins," said Pommier. "Our findings indicate that PARP inhibitors should be categorized according to their potency to trap PARP, in addition to their ability to inhibit DNA repair. This is important because it might explain differences in the results of clinical trials using distinct PARP inhibitors."
In further experiments, the researchers identified several genetic mutations in post-replication repair and Fanconi anemia pathways that, like BRCA1 and BRCA2 mutations, sensitized cultured cells to the toxic effects of trapped PARP protein-DNA complexes.
"These data suggest that patients with cancers deficient in these PARP inhibitor-sensitizing genes might benefit from treatment with PARP inhibitors," said Pommier. "It is clear, however, that this hypothesis will require rigorous testing before being broadly translated to the clinic."