Valuable tool for predicting pain genes in people
Scientists in Australia and Austria have described a "network map" of genes involved in pain perception. The work, published in the journal PLOS Genetics should help identify new analgesic drugs.
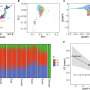
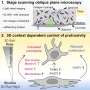
Dr Greg Neely from the Garvan institute of Medical Research in Sydney and Professor Josef Penninger from the Austrian Academy of Sciences in Vienna had previously screened the 14,000 genes in the fruit fly genome and identified 580 genes associated with heat perception. In the current study, using a database from the US National Centre for Biotechnology Information, they noted roughly 400 equivalent genes in people, 35% of which are already suspected to be pain genes.
The map they constructed using fly and human data includes many known genes, as well as hundreds of new genes and pathways, and demonstrates exceptional evolutionary conservation of molecular mechanisms across species. This should not be surprising, as every creature must be able to identify a source of pain or danger in order to survive.
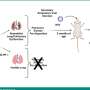
Comparing fly with human data, they could see that a particular kind of molecular signaling (phospholipid signaling), already implicated in pain processing, appeared in the pain network. Further, they demonstrated the importance of two enzymes that make phospholipids, by removing those enzymes from mice, making them hypersensitive to heat pain.
"Pain affects hundreds of millions of people, and is a research field badly in need of new approaches and discoveries," said Dr Neely.
"The fact that evolution has done such a remarkable job of conserving pain genes across species makes our fly data very useful, because much of it translates to rodents and people.
"We are able to test our hypotheses in mice, and if a gene or pathway or process functions as we predict, there is a good chance it will also apply to people.
"By cross-referencing fly data with human information already in the public domain – like gene expression profiling or genetic association studies – we know we'll be able to pinpoint new therapeutic targets."
More information: PLOS Genetics. doi:10.1371/journal.pgen.1003071