March 28, 2013 report
Research team uncovers shape of transmembrane protein partly responsible for antibiotic resistance
(Medical Xpress)—Researchers from the University of Tokyo have uncovered the physical layout of a transmembrane protein that the tiny organism Archaea relies on to keep toxins out of its cells. The protein, the team explains in their paper published in the journal Nature, is also believed to be responsible for preventing the porting of therapeutic drugs into the cell body, stopping their usefulness.
Transmembrane proteins are members of a type of protein family called multidrug and toxic compound extrusion transporters (MATEs)—they're located in the membranes of cells and their purpose is to protect the cell from intrusion by toxic materials. To do so, they carry only molecules identified as safe through the membrane into the inner portion of individual cells. Researchers are keen to gain a better understanding of MATEs because they are believed to also sometimes block therapeutic drugs that are meant to enter cells to help fight off infections or some cancers. In this new effort, the research team has mapped the configuration of a MATE in an Archaea, which the team believes will help in doing the same for cells in humans.
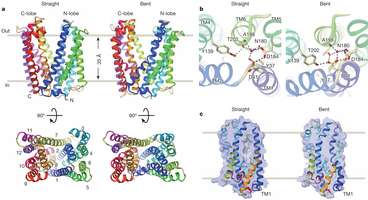
Specifically, the team found that the MATEs under study were shaped like the letter "V" and were positioned with the open end facing outwards. They believe that when a toxic molecule moves into the valley of the "V," one of the straight walls bends outward, pushing the toxin away. This is the third mapping of a MATE—prior research by other teams has found the configuration of two other bacteria cells.
The research team also discovered a certain peptide that was able to get in the way of the MATE doing its job, possibly leading to the creation of a drug that can block MATE activity long enough to allow beneficial drugs to enter cells in need of help. The team points out that the peptide in their study hasn't been found to be useful in humans, but its discovery leads to hope of finding one that will, which in the long run would help patients' cells accept beneficial drugs. Once that happens, they add, it will still have to undergo extensive testing to ensure it doesn't also cause harmful side effects, such as triggering an immune response.
More information: Structural basis for the drug extrusion mechanism by a MATE multidrug transporter, Nature (2013) doi:10.1038/nature12014 . www.nature.com/nature/journal/ … ull/nature12014.html
Abstract
Multidrug and toxic compound extrusion (MATE) family transporters are conserved in the three primary domains of life (Archaea, Bacteria and Eukarya), and export xenobiotics using an electrochemical gradient of H+ or Na+ across the membrane. MATE transporters confer multidrug resistance to bacterial pathogens and cancer cells7, thus causing critical reductions in the therapeutic efficacies of antibiotics and anti-cancer drugs, respectively. Therefore, the development of MATE inhibitors has long been awaited in the field of clinical medicine. Here we present the crystal structures of the H+-driven MATE transporter from Pyrococcus furiosus in two distinct apo-form conformations, and in complexes with a derivative of the antibacterial drug norfloxacin and three in vitro selected thioether-macrocyclic peptides, at 2.1–3.0 Å resolutions. The structures, combined with functional analyses, show that the protonation of Asp 41 on the amino (N)-terminal lobe induces the bending of TM1, which in turn collapses the N-lobe cavity, thereby extruding the substrate drug to the extracellular space. Moreover, the macrocyclic peptides bind the central cleft in distinct manners, which correlate with their inhibitory activities. The strongest inhibitory peptide that occupies the N-lobe cavity may pave the way towards the development of efficient inhibitors against MATE transporters.
© 2013 Medical Xpress