Insight into cell survival
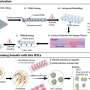
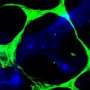
![Catalytic centre in Atg3. In the absence of Atg12-Atg5, a catalytic Cys234 (magenta) is oriented away from Thr213 (orange) which is essential for Atg3 activity. When Atg3 interacts with Atg12-Atg5, Atg3 causes a conformational change, which reorients Cys234 toward Thr213 and mediates the formation of a Atg8-phosphatidylethanolamine. Tokyo Institute of Technology research: An insight into cell survival [research]](https://scx1.b-cdn.net/csz/news/800a/2013/tokyoinstitu.jpg)
Researchers at Tokyo Institute of Technology report details on the biological mechanisms through which cells degrade own cellular material, allowing them to survive starvation conditions.
Protein conformation studies by a collaboration of researchers in Japan identify changes in protein shape that contribute to autophagy, a major system for degrading cellular material and maintaining cells. The results clarify stages in autophagy that were previously a mystery.
"This study reveals the mechanism of the key reaction that drives membrane biogenesis during autophagy," explain Yoshinori Ohsumi, Machiko Sakoh-Nakatogawa and colleagues at Tokyo Institute of Technology, Institute of Microbial Chemistry and Hokkaido University.
Autophagy is a bulk degradation system essential for cell survival under starvation conditions, and also allows cells to control over the quality of cell components. It is known to be linked to neurodegenerative and hepatic diseases and the subject of intense research interest.
Cell degradation takes place through the formation of double-membraned vesicles —'autophagasomes'—which engulf the cytosolic components. Previous studies have identified a number of autophagy-related (Atg) proteins linked to the biogenesis of the autophagasome.
Similarities have been drawn between the catalytic roles played by some of these Atg proteins and enzymes in other similar processes. However the Atg proteins differ structurally from other known enzymes with similar roles, so that how they activated different stages in autophagy was as the researchers point out "totally unknown".
Ohsumi, Sakoh-Nakatogawa and colleagues used biochemical assays, X ray crystallography and mutational analyses to study the conformation of the proteins and how their activities changed for different mutant variants of the proteins. The results determine configuration of amino acid residues on the proteins that alter the protein conformation and activate specific stages in autophagy.
The researchers also identify a regulatory function for this stage in the mechanism, as it ensures the proteins activate specifically at autophagy-related membranes.
Autophagy
Autophagy is essential for cell survival under starvation condition supplying nutrient source derived from degradation of cellular materials. Autophagy also plays a housekeeping role in removing unwanted or dysfunctional cytosolic components and pathogens. .
Authophagy-related (Atg) proteins and the ubiquitination enzymes E1, E2 and E3
There are 18 Atg proteins that have been linked with autophagy processes. These proteins have different amino acid residues at various locations on the protein structure that contribute to catalysing the steps involved in autophagy. The researchers studied mutant proteins with different variants of these residues to clarify the role the residues play in autophagy.
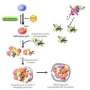
Previous studies have drawn similarities between some of these Atg proteins in the series of reactions they undergo and protein ubiquitination, the modification of a protein by another so-called ubiquitin protein. The name ubiquitin originates from the ubiquity of the protein as it is found in almost all organisms and directs proteins to degradation by the proteasome or to transport to different compartments and locations in the cell.
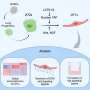
Three stages have been identified in the process of ubiquitination: activation of ubitquitin by an enzyme E1, transfer of ubiquitin from E1 to a ubiquitin-conjugating enzyme E2 and the ubiquitination cascade, using one of hundreds of identified 'E3' ubiquitin-protein ligases. In authophagy Atg7 has been described as an E1 enzyme and Atg3 and Atg10 as E2 enzymes. In previous work the researchers reported interaction between the Atg12-Atg5 conjugate and Atg3, indicating that Atg12-Atg5 acts as an E3 enzyme. However as the conjugate lacked domains typical of E3 enzymes, how Atg12–Atg5 activates Atg3 had remained a mystery until the report of these latest results.
Experimental approaches
X ray crystallography provided details of the Atg protein structures. Studies in vitro complemented this data to identify the protein conformations.
Based on the structural information, the researchers introduced Cys residue at some locations on Atg3 and examined an intramolecular disulphide bond formation between the introduced Cys residue and a catalytic Cys residue which indicates the proximity of these Cys residues. From the intramolecular disulphide bond formation with or without an interaction of Atg12-Atg5, the researchers identified that Atg12-Atg5 introduces a conformational change in the catalytic centre of Atg3. Effects of various mutations for an intramolecular disulphide bond formation in the Cys mutants of Atg3 allowed the researchers to identify the role of the mutated amino acid residues in the suppression of activity in the absence of Atg12-Atg5.
Atg12-Atg5 activation of Atg3 regulatory mechanism
From the biochemical assays, mutational analyses and conformational data the researchers determined a structural change induced as Atg12-Atg5 binds to Atg3. The structural change activates the catalytic centre of Atg3.
This stage has a regulatory function in the mechanism. Atg3 mediates the formation of a Atg8– phosphatidylethanolamine conjugate, which localizes to form autophagosomal membranes. Atg3 may be activated by Atg12–Atg5 at the preautophagosomal structure to specifically produce Atg8– phosphatidylethanolamine on autophagy-related membranes.
More information: Sakoh-Nakatogawa, M. et al. Atg12–Atg5 conjugate enhances E2 activity of Atg3 by rearranging its catalytic site. Nature Structural and Molecular Biology 20, 433-439 (2013). doi: 10.1038/nsmb.2527