New study to give insight into the public health risks of antibiotic resistant bacteria
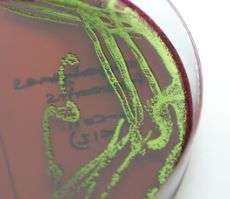
Scientists at Queen Mary, University of London are part of a national study seeking to establish the most significant reservoirs of an antibiotic resistant bacteria known as ESBL-positive E.coli that cause human and animal disease.
The findings of this study will help to develop intervention strategies aimed at reducing the numbers of urinary tract and bloodstream infections caused by these bacteria.
The research is being led by Public Health England, with key collaborators including Queen Mary, the Animal Health and Veterinary Laboratories Agency, The University of Cardiff, The University of East Anglia, The University of Glasgow and Health Protection Scotland.
The study will look at sewage, farm slurry and raw meat to determine whether there are any potential risks to human health from these reservoirs. It will also look for the presence of ESBL positive E. coli in routinely collected stool samples to assess the proportion of people carrying it in their gut who have no symptoms of illness. Genetic studies will be used to compare the resistant bacteria from the various sources to identify features that may contribute to how easily they spread and their ability to cause disease.
E.coli is a bacterium that lives in the guts of humans and many other animals. Colonisation of the gut by E. coli is perfectly normal and harmless, although some types can cause diarrhoea. However, E.coli is also the commonest cause of urinary tract and bloodstream infections, which usually require antibiotic treatment.
Not all types of ESBL-positive E.coli bacteria cause human disease, and the contribution to human disease made by resistant strains from animals, meat and environmental sources is not well understood.
Resistant strains of E.coli are an increasing problem, reducing the number of antibiotics that a doctor can use for treatment. Many of the resistant strains produce enzymes called ESBLs (Extended-Spectrum Beta-Lactamases), which make them resistant to most penicillin-like antibiotics. E.coli with ESBLs can also be found in food animals, raw retail meat, sewage and river water, but whether these reservoirs pose any public health risk is poorly understood.
Dr David Wareham, a consultant microbiologist and clinical senior lecturer at the Blizard Institute at Barts and The London School of Medicine and Dentistry, Queen Mary, said: "The issue of antibiotic resistant E.coli is certainly moving up the risk register in hospitals. It is having a big impact on prescribing of antibiotics and is leading to more broad-based antibiotics being prescribed which in turn is driving up the problem.
"Due to our urban location and diverse local population, the selection of Queen Mary as one of the regional centres will help us define the impact of travel and migration on the spread of resistance, as well as having immediate benefits for the local community."
Professor Neil Woodford, Head of the Antimicrobial Resistance and Healthcare Associated Infections Reference Unit at PHE and Honorary Professor of Microbiology at Queen Mary, said: "The risks posed to human health by resistant E.coli from non-human reservoirs are not fully understood. This study will help to disentangle this complex interrelationship.
"Treatment of infections caused by resistant E. coli can be difficult, which is why we need to understand the risks better. Having said that, we want to reassure the public that presence of these bacteria in the gut does not require antibiotic treatment and is usually temporary. Most colonized people never develop an infection caused by the resistant strain.
"This study is very important because its results will help to shape future intervention strategies to reduce the spread of these antibiotic-resistant strains of bacteria and to reduce the numbers of infections that they cause."
The study is funded by the Department of Health for three years.