Prediction of the future flu virus
Every year, influenza outbreaks claim hundreds of thousands of human lives. Though vaccination against flu is fairly efficient, the disease is difficult to exterminate because of the high evolutionary rate of the flu virus. Every year, new flu strains spread over the planet that differ slightly from those that were common a year before, which helps the virus to escape the immune response, and possibly compromises the efficiency of anti-viral drugs. Furthermore, from time to time, a drastically new strain appears, posing a threat of human pandemic. Both processes are due to changes in the viral genome, but of a different degree.
The differences in the seasonal flu usually result from point mutations in the influenza virus genes, while major pandemics are often connected to profound genetic shifts known as reassortments. The link between these two phenomena was studied for the first time by the Russian research team from the Faculty of Bioengineering and Bioinformatics of the Moscow State University (MSU) in collaboration with the Central Research Institute of Epidemiology and the Institute for Information Transmission Problems of the Russian Academy of Sciences (IITP). One of the authors, professor Alexey Kondrashov, is also affiliated with the University of Michigan. The results were published on January 9 in PLoS Genetics.
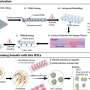
Georgii Bazykin, the corresponding author on the paper, who is a leading researcher at the Faculty of Bioengineering and Bioinformatics at the MSU and the head of the Molecular Evolution division at IITP, explains: "Influenza virus genome consists not of a single DNA or RNA molecule as many other viruses do, but of eight individual segments resembling in some sense the chromosomes of the human genome. Every segment is a separate RNA molecule." If different strains co-infect a single cell, their genomes may exchange these segments in a process called reassortment. This may lead to emergence of a novel genome consisting, for instance, of three segments obtained from one viral genome and five segments from another.
Bazykin says, "Most major flu epidemics that we know were caused by such reassortments. When you analyze the strains that have caused these outbreaks, you find that they had combinations of viral genome segments that were never seen together before. This was the case for the 1957 and the 1968 pandemics, as well as for the swine flu in 2009. The deadliest Spanish flu pandemic of 1918 had probably the same nature, although it is hard to be certain for the events so distant in time."
A reassortment may produce a highly virulent strain, because a strong genetic shift makes it "unfamiliar" to the immune system of most humans, which allows the virus to spread efficiently throughout the population.
This is the evolutionary scenario known as antigenic shift. Another path, known as antigenic drift, is a process of gradual accumulation of smaller mutations. These mutations cause changes in the viral antigenic proteins, primarily, the surface antigens hemagglutinin (HA) and neuraminidase (NA). The genes coding for these proteins evolve rapidly in the course of the arms race between the virus and immune system.
"The seasonal flu outbreaks are primarily caused by this antigenic drift,—explains Georgii Bazykin.—Hence every year many of us catch a flu caused by a new strain of the constantly evolving virus."
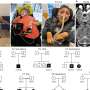
The relationship between these two processes,—antigenic shift and antigenic drift,—has never been studied before. To fill this gap, Russian researchers first aimed on localizing the reassortments that the virus had experienced during its evolution. They studied the Influenza A H3N2 virus that entered the human population in 1968. Using a data base containing 1376 sequenced viral genomes, they used bioinformatic techniques to reconstruct the history of reassortments, and to pinpoint the reassortments on the evolutionary tree of the virus. They then checked the hypothesis that genetic shift causes subsequent genetic drift; i.e., that reassortments increase the frequency of smaller, point mutations that appear when individual nucleotides are replaced. The hypothesis was supported by the data: indeed, a reassortment increases the subsequent rate of single-nucleotide substitutions.
"We believe that this effect is connected to the fact that reassorted genes have to operate in a new genetic environment," says Bazykin. "Since genes are connected to each other, if gene A has changed, a new version of gene B is also likely to be preferable. As a result, every reassortment event is followed by a trail of additional point mutations."
Since reassortments produce the most virulent pandemic-causing strains, the results of this work may be relevant to prediction of the future emergence of such potentially dangerous outbreaks.