Using computer modeling to understand how messy cells contribute to cancer
Life can be messy at all scales, requiring different organizational strategies—from cleaning the house, to removing damaged or expired cells from the body to avoid cancer progression.
In a messy house, people use computers to manage paper and photo clutter; companies use computer systems to track their inventory. Now a team of researchers at Vanderbilt University in Nashville, Tenn., is taking a similar approach to cell-molecular inventory control for cancer. They have created computer models, using their programming framework (PySB), which enable them to explore the complex biochemical processes that drive cancer growth.
"Our hypothesis is that understanding how the cell uses their protein inventory will lead to understanding why cells dysregulate and become carcinogenic. We expect model outputs will lead to novel, targeted cancer therapies—possibly by 2019," explained researcher Carlos F. Lopez, who will present the work at the 58th annual Biophysical Society Meeting in San Francisco, Feb.15-19.
Lopez is interested in understanding how cells in multicellular organisms engage programmed cell death—so-called "cell suicide"—for cellular removal. It is a natural part of many cells' life cycle.
When cancer cells avoid programmed cell death, uncontrolled growth fuels tumor progression. The Vanderbilt team expects their computer models to identify what goes wrong in these cases, at a speed and scale never before possible. Lopez noted: "We are bridging the nanoscale molecular-level biochemical interactions with the macroscale cancer tumor outcomes, which is a huge range in scales. Most people don't realize this, but molecular chemical reactions at the nanometer and nanosecond level affect things that happen at the timescale level of years—nine orders of magnitude in space and time! For comparison, a nanosecond is to a second like a second is to one century."
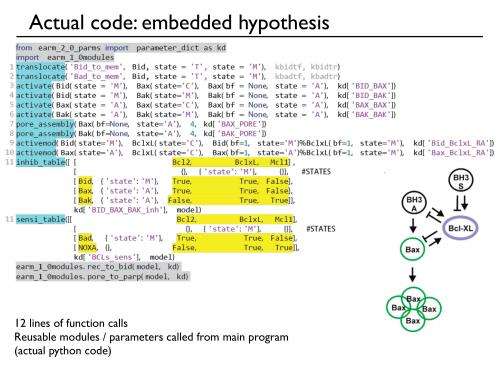
Rather than listing the cellular biochemical reactions by hand, PySB enables the researchers to "write" the biochemical cellular processes as computer programs. "With this approach we can create, simulate, and explore multiple complex mathematical systems that represent biology with ease," Dr. Lopez said.
The models aim to understand how healthy cells normally self-destruct when removal is needed, and why cancer cells avoid the removal signals. If the approach is validated over the next few years, it could lead to the use of predictive mathematical models, representing biochemical signaling processes, to forecast how a cell, group of cells, or a tumor, would respond to single or multiple drug combinations. Targeted, individualized and optimized cancer treatments could then be tailored to each patient.
More information: The presentation "PySB: A Modeling Framework to Explore Biochemical Signaling Processes and Cell-decisions" by Carlos F. Lopez and Shawn P. Garbett will be at 1:15 p.m. on Wednesday, February 19, 2014 in Room 304 in San Francisco's Moscone Convention Center. Abstract: tinyurl.com/n2nw99a