DNA origami nano-tool provides important clue to cancer
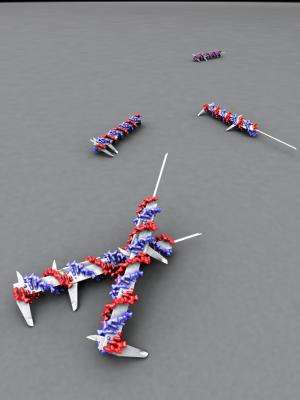

Researchers at Karolinska Institutet have headed a study that provides new knowledge about the EphA2 receptor, which is significant in several forms of cancer. This is important knowledge in itself – but just as important is how this study, which is published in the highly respected journal Nature Methods, was conducted. The researchers used the method of DNA origami, in which a DNA molecule is shaped into a nanostructure, and used these structures to test theories about cell signalling.
It was previously known that the EphA2 receptor played a part in several forms of cancer, such as breast cancer. The ligand, i.e., the protein that communicates with the receptor, is known as an ephrin molecule. Researchers have had a hypothesis that the distance between different ligands – in this case the distance between ephrin molecules – affects the level of activity in the communicating receptor of the adjacent cells.
The Swedish researchers set out to test this hypothesis. They used DNA building blocks to form a stable rod. This has then been used as a very accurate measure of the distance between molecules.
"We use DNA as the construction material for a tool that we can experiment with", says Björn Högberg, principal investigator at the Department of Neuroscience. "The genetic code of the DNA in these structures is less important in this case."
The researchers attached proteins, or ephrins, to the DNA rod at various intervals, for example 40 or 100 nanometres apart. The DNA rods were then placed in a solution containing breast cancer cells. In the next step, the researchers looked at how active EphA2 was in these cancer cells.
Placed close together
It turned out that if the ephrin molecules were placed close together on the DNA rod, the receptor in question became more active in the cancer cells, and the cells also became less invasive in respect of the surrounding cells, which could be an indication that they became less prone to metastasis. This was true even though the amount of protein was the same throughout the experiments, i.e., the number of attached molecules remained the same.

"For the very first time, we have been able to prove this hypothesis: the activity of EphA2 is influenced by how closely spaced the ligands are on the surrounding cells", says Björn Högberg. "This is an important result in itself, but the point of our study is also that we have developed a method for examining how cells react to nearby cells in a controlled environment, using our custom DNA nano-calipers."
The researchers describe the cell communication as a form of Braille, where the cells somehow sense the protein patterns of nearby cells, and where the important thing is not only the amount of proteins, but to a great extent the distance between them as well. This study found that a cluster of proteins would communicate more actively than sparsely spaced proteins, even if the concentration was the same.
"This is a model that can help us learn more about the importance of the spatial organization of proteins in the cell membrane to how cells communicate with each other, something that will hopefully pave the way for a brand new approach to pharmaceuticals in the long term", says Ana Teixeira, a principal investigator at the Department of Cell and Molecular Biology. "Today, the function of the pharmaceuticals is often to completely block proteins or receptors, but it is possible that we should rather look at the proteins in their biological context, where the clustering and placement of various proteins are relevant factors for the effect of a drug. This is probably an area where there is important knowledge to obtain, and this is a way of doing it."
More information: Spatial Control of Membrane Receptor Function using Ligand Nano–Calipers. Alan Shaw, Vanessa Lundin, Ekaterina Petrova, Ferenc Fördős, Erik Benson, Abdullah Al–Amin, Anna Herland, Andries Blokzijl, Björn Högberg och Ana I. Teixeira. Nature Methods, online July 6 2014; DOI: 10.1038/nmeth.3025