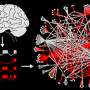
Largest-ever map of the human interactome predicts new cancer genes
Scientists have created the largest-scale map to date of direct interactions between proteins encoded by the human genome and newly predicted dozens of genes to be involved in cancer.
The new "human interactome" map describes about 14,000 direct interactions between proteins. The interactome is the network formed by proteins and other cellular components that 'stick together.' The new map is over four times larger than any previous map of its kind, containing more high-quality interactions than have come from all previous studies put together.
CIFAR Senior Fellow Frederick Roth, along with Marc Vidal (Dana-Farber Cancer Institute and Harvard Medical School) co-led the international research team that wrote the forthcoming Nov. 20 paper in Cell. Roth is a senior investigator at Mount Sinai Hospital's Lunenfeld-Tanenbaum Research Institute and a professor at the Donnelly Centre for Cellular and Biomolecular Research, University of Toronto. He is also a Canada Excellence Research Chair in Integrative Biology.
Using lab experiments, the scientists identified interactions and then, using computer modelling, they zoomed in on proteins that 'connect' to one or more other cancer proteins.
"We show, really for the first time, that cancer proteins are more likely to interconnect with one another than they are to connect to randomly chosen non-cancer proteins," Roth says.
"Once you see that proteins associated to the same disease are more likely to connect to each other, now you can use this network of interactions as a prediction tool to find new cancer proteins, and the genes they encode," says Roth. For example, two known cancer genes encoded two proteins that interacted with CTBP2, a protein encoded at a location tied to prostate cancer, which can spread to nearby lymph nodes. These two proteins are implicated in lymphoid tumours, suggesting that CTBP2 plays a role in the development of lymphoid tumours.
Using their predictive method, the researchers found that 60 of their predicted cancer genes fit into a known cancer pathway.
Discoveries like these are crucial for understanding how cancer and other diseases develop and ultimately, how to treat and prevent them. The vast majority of protein interactions in the human body are a mystery. Roth likens a doctor asked to treat a patient's disease to an auto mechanic.
"How can we ask someone to fix a car with an incomplete list of parts and no guidance on how the parts fit together?" he says.
Each gene can encode multiple parts, and researchers are working toward a comprehensive understanding of what all those parts are, where they are found within the cells of our bodies, and how they connect to each other. Studies in baker's yeast have mapped interactions at genome scale, but the new study is the first to approach this scale in humans.
The study also reveals that the network of protein interactions in humans covers a much broader range of genes than some past research has suggested. Studies often focus on 'popular' proteins that are already known to be tied to disease or to be interesting for other reasons, which has created a bias in our understanding of interactions, Roth says.
"One major conclusion of the paper is that when you look systematically for interactions, you find them everywhere," he says.
Roth says the research is central to the goals of CIFAR's Genetic Networks program, particularly building the map of how an organism's genotype, the set of its genes, connects to an organism's phenotype, its characteristics that include appearance and predisposition to disease. Knowledge of interactions is likely to inform worldwide efforts to sequence and interpret cancer genomes.