February 10, 2015 feature
Of cancer and chaos: Single base mutation induces cancer-like gene profile and major unexpected impact on phenotype
(Medical Xpress)—In chaos theory – the study of dynamical systems highly sensitive to initial conditions – was first described in 1890 by Henri Poincaré1, followed by work by Jacques Hadamard in 1898 and Pierre Duhem in 1908. Then, in his 1963 paper2, meteorologist Edward N. Lorenz theoretically demonstrated how in deterministic nonlinear systems, slightly differing initial states can evolve into very different later states – an effect commonly known as the butterfly effect. (While Lorenz did not use the term in his paper – it was previously introduced by Ray Bradbury in his 1952 science fiction story, A Sound of Thunder – it is attributed to him: the title of his talk at the 139th meeting of the American Association for the Advancement of Science in 1972, Does the flap of a butterfly's wings in Brazil set off a tornado in Texas?, was anecdotally coined by fellow meteorologist Philip Merilees.) That said, while chaotic behavior in many natural systems can be studied through analysis of a chaotic mathematical model or analytical techniques such as recurrence plots and Poincaré maps, there is not universal agreement that weather systems are so exquisitely sensitive to initial conditions, with alternative explanations including model error and oversimplified Lorenz equations.
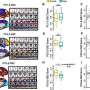
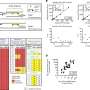
Recently, scientists at The Scripps Research Institute published a paper reporting that a single base substitution in one allele of a gene in a human breast epithelial cell not only leads to a gene expression profile similar to that of basal breast cancer, but also causes extensive remodeling of gene signatures that are not known to be connected to the activity of that gene. In other words, they say, "a cancer-specific mutation…has an unexpectedly deep and broad impact on the phenotypic properties of the cell." In their paper, they note that "the butterfly concept applies perfectly to the single base substitution studied….It is the smallest genetic change that can be introduced in a cell, yet it has immense consequences." The researchers observed that the magnitude and extent of changes seen in the mutated cells suggest that the cell line modification "has inadvertently also selected for other mutations, amplifications, or deletions that could be responsible for some of the changes observed." In general terms, they say, this suggests that the effects of a somatic mutation are determined by the cell type in which the mutation occurs.
Prof. Peter K. Vogt discussed the paper that he, Dr. Jonathan R. Hart and their co-authors published in Proceedings of the National Academy of Sciences. "Comparing the exome, transcriptome, proteome and metabolome of two cell lines can be done with current technology," Vogt tells Medical Xpress. (The exome is the part of the genome that consists of all DNA transcribed into mature RNA in cells of any type; the transcriptome comprises all RNA molecules found in a specified cell population – including mRNA, rRNA, tRNA, and other non-coding RNA; the proteome is the entire set of proteins expressed by a genome, cell, tissue or organism at a certain time under defined conditions; and the metabolome refers to all biological reaction products found within a biological sample, such as a cell, a cellular organelle, an organ, a tissue, a tissue extract, a biofluid, or an entire organism.)
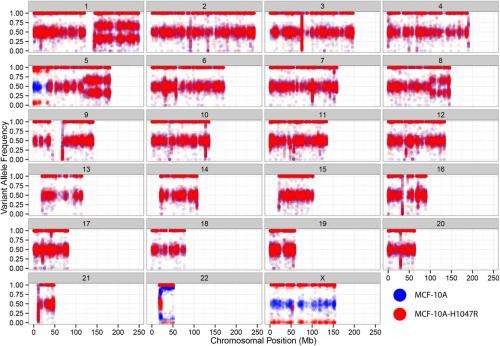
"However," Vogt continues, "for such comparisons, the important step is showing that the single mutation is the only relevant difference between the cells." Specifically, he notes, in the case of MCF-10A and MCF-10A-H1047R cells, there are other genetic differences besides the H1047R mutation – and while these differences do not include cancer-relevant genes, in a strictly molecular sense the cell lines are not isogenic (that is, characterized by essentially identical genes).
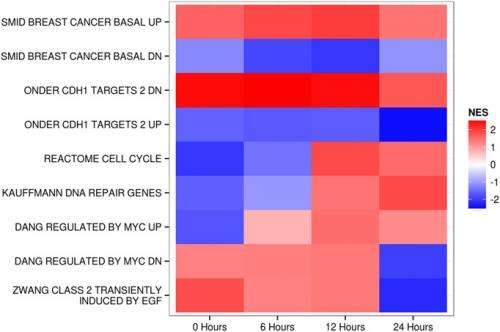
The scientists derived two key insights from this study. "The smallest possible genetic change in PI3K – a known driver gene in cancer – can lead to an extensive remodeling of the cell that reaches far beyond the established activities and roles that can be attributed to PI3K," Vogt points out. "In addition, in the case of PI3K, a gain-of-function mutation drives the proteome and transcriptome of a breast epithelial cell line to show a gene signature that closely resembles that of mature basal breast cancer, but not that of other histological types of breast cancer. It therefore appears that MCF-10A cells are pre-programmed to respond to PI3K mutation by activating a definitive pattern of phenotypic changes." To the researchers, this means that future work could thus focus on the differences between the H1047R mutant-induced changes and those recurrently appearing in basal breast cancer.
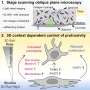
"Our observations also raise several questions," Vogt adds. "First, to what extent are the mutant-induced changes in the proteome and transcriptome continuously dependent on the gain of function in PI3K? To answer this question, we've used small molecule inhibitors that act downstream of PI3K." This step can revert some but not all mutant-associated changes in MCF-10A-H1047R cells, suggesting that some of the changes may be irreversible, possibly by epigenetic mechanisms – that is, long-term changes in a cell's transcriptional potential, gene expression or cellular phenotype not caused by changes in the DNA sequence.
The second question concerns the state of differentiation of the MCF-10A cells and the effect of this differentiated state on the response to a single point mutation. "We expect that the same mutant introduced into another cell type – for example, an intestinal epithelial cell – will generate a gene signature that unrelated to that of basal breast cancer, and may therefore resemble that of another type of cancer."
Interestingly, the scientists found that the mutant cell system has several key features in common with the butterfly effect. "Firstly," Vogt explains, "we're dealing with a very small initiating change that has huge downstream consequences. Secondly, this change acts on a system that is complex and nonlinear. While cells are dynamic adaptive systems, adaptability is restricted by genetic and epigenetic information – for example, a cell's lineage and state of differentiation. "In contrast to the original formulation of the butterfly effect in fluid flow, we're only just beginning to unravel the cell's complex dynamics."
The scientists state that their work opens the way to several future investigations, including exploration of the epigenetic mechanisms that convey mutant-independence on some of the phenotypic changes that they have identified. "Due to their frequent involvement in chromatin remodeling" – the dynamic modification of chromatin architecture to allow access of condensed genomic DNA to the regulatory transcription machinery proteins, and thereby control gene expression – "epigenetic changes bring up the broad area of non-coding RNAs," Vogt says. "Our RNAseq data include information on the non-coding transcriptome and will be used for an analysis of epigenetics." RNASeq, or RNA Sequencing, achieves transcriptome profiling by using next-generation sequencing to determine genomic RNA presence and quantity a given moment in time.
"Moving in another direction," Vogt tells Medical Xpress, "generating an isogenic pair of intestinal epithelial cells with the single H1047R mutation will allow an assessment of the effect of cell differentiation on cancer gene signatures. In addition, more recent techniques such as CRISPR/Cas9 would allow us to insert such a modification in a less disruptive fashion than has been historically possible." CRISPR/Cas9 (Clustered Regularly Interspaced Short Palindromic Repeats – DNA loci containing short repetitions of base sequences – and CRISPR associated protein 9 refers to a genome engineering/editing tool that can also create programmable transcription factors, allowing scientists to target and activate or silence specific genes – and that may be capable of editing the genomes of entire populations of organisms through the fabrication of Cas9-based endonuclease gene drives. (Endonucleases are enzymes that cleave the phosphodiester bond within a polynucleotide chain.)
Vogt also mentions a complementary study in which it will be important to determine the effect of other cancer mutations, such as those in the RAS oncogene on cellular remodeling. (An oncogene has the potential to cause cancer by causing cells designated for apoptosis, or rapid cell death, to survive and proliferate instead.)
"Finally," Vogt concludes, "today's gene engineering technology will allow the generation of numerous isogenic cell lines that differ in a variety of single genes – coding as well as non-coding – making it possible to examine the butterfly effect in these new settings."
More information: The butterfly effect in cancer: A single base mutation can remodel the cell, Proceedings of the National Academy of Sciences (2015) 112(4):1131-1136, doi:10.1073/pnas.1424012112
Related:
1Sur les intégrales de fonctions a multiplicateurs et leur application au développement des fonctions abéliennes en séries trigonométriques, Acta Mathematica (1890) 13(2):4-174, doi:10.1007/BF02981629
2Deterministic Nonperiodic Flow, Journal of the Atmospheric Sciences (1963) 20:130–141, doi:10.1175/1520-0469(1963)020<0130:DNF>2.0.CO;2 (PDF)
© 2015 Medical Xpress