Researchers discover new clues for treatment of antibiotic-resistant bacteria
Researchers at the University of Georgia have identified a previously unknown process that many bacteria, including those that cause disease in humans, use to survive. Their discovery could lead to new therapies for bacterial infections like MRSA and tuberculosis that are resistant to current antibiotic treatments.
Their discovery centers on the ways in which bacteria use an iron compound known as heme. Most commonly recognized as a fundamental component of hemoglobin—the red pigment in blood responsible for the transportation of oxygen—heme also provides some of the essential nutrition that bacteria require to grow within a host.
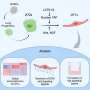
"We found an enzyme that only gram-positive bacteria have, and they use this enzyme in a unique way to make heme," said Harry Dailey, professor of microbiology and lead author of the paper describing their discovery in the Proceedings of the National Academy of Sciences. "Heme is essential for these organisms; they have to make heme, so if we can find a way to disrupt this process, the bacteria would die."
More than 2 million people become infected with bacteria that are resistant to antibiotics each year in the U.S., and at least 23,000 people die as a direct result of these infections, according to the Centers for Disease Control and Prevention.
The enzyme discovered in Dailey's lab could serve as an attractive target for the next generation of antibacterial drugs, and his laboratory is partnered with other institutions to explore the development of new therapeutics.
"One of the great things about this enzyme in terms of treatment is that it is only found in gram positive bacteria-it's not found anywhere else," said Dailey, who is also director of UGA's Biomedical and Health Sciences Institute. "So if you target the enzyme with a therapeutic, you won't damage any of the other essential mechanisms that we need to remain healthy."
Their discovery also challenges some of the established scientific literature related to the production of heme, which has remained largely unchanged for decades.
"We are obviously very excited about the potential medical applications related to this enzyme, but our experiments also tell us a lot about the evolutionary history driving the behavior of organisms we study every day," Dailey said. "We have assumed for more than 50 years that we understood these pathways well, but it may not be as clear cut as we once thought."
More information: "Noncanonical coproporphyrin-dependent bacterial heme biosynthesis pathway that does not use protoporphyrin." PNAS 2015 112 (7) 2210-2215; published ahead of print February 2, 2015, DOI: 10.1073/pnas.1416285112