'Most comprehensive map' of human epigenomes is unveiled
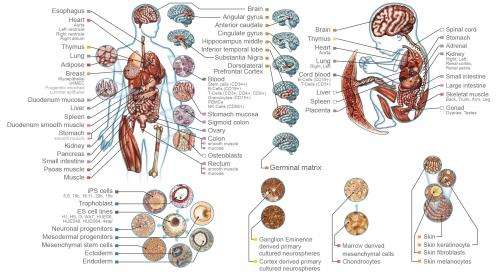
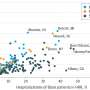
Two dozen scientific papers published online simultaneously on Feb. 18, 2015 present the first comprehensive maps and analyses of the epigenomes of a wide array of human cell and tissue types. Epigenomes are patterns of chemical annotations to the genome that determine whether, how, and when genes are activated.
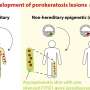
Because epigenomes orchestrate normal development of the body, and disruptions in epigenetic control are known to be involved in a wide range of disorders from cancer to autism to heart disease, the massive trove of data is expected to yield many new insights into human biology in both health and disease.
The 24 papers describing human epigenomes will appear in print on Feb. 19, 2015 in the journal Nature and in six other journals under the aegis of Nature Publishing Group. Collectively, the papers are a culmination of years of research by hundreds of participants in the Roadmap Epigenomics Program (REP), first proposed in 2006 by academic scientists and key members of the National Institutes of Health. All will be freely available at Nature's Epigenome Roadmap website.
"The DNA sequence of the human genome is identical in all cells of the body, but cell types—such as heart, brain or skin cells—have unique characteristics and are uniquely susceptible to various diseases," said UC San Francisco's Joseph F. Costello, PhD, director of one of four NIH Roadmap Epigenome Mapping Centers (REMC) that contributed data to the REP. "By guiding how genes are expressed, epigenomes allow cells carrying the same DNA to differentiate into the more than 200 types found in the human body."
In cancer research, said Costello, the new data will hasten a merging of genomic and epigenomic perspectives that was already underway. "You've had cancer researchers studying the genome—the role of mutations, deletions, and so on—and others studying epigenomes. They've almost been working on parallel tracks, and they didn't talk to each other all that much. Over the past five or six years, there's been a reframing of the discussion, because the most recurrent mutations in cancer affect epigenomic regulators. So the way mutations in the genome play out is through epigenomic mechanisms, and major pharmaceutical companies now view epigenomes as an important target."
Costello holds the Karen Osney Brownstein Endowed Chair in Neuro-Oncology in the UCSF Department of Neurological Surgery, and is a member of the UCSF Helen Diller Family Comprehensive Cancer Center (HDFCCC).
The overarching findings of the REP, which include data on 111 distinct human epigenomes from all four REMCs as well as from dozens of individual labs around the world, are covered in a Nature paper for which Manolis Kellis, PhD, of Massachusetts Institute of Technology (MIT) and the Broad Institute of MIT and Harvard, is senior author. In addition to the many implications for normal human biology of "the most comprehensive map of the human epigenomic landscape so far," the authors write, "our data sets will be valuable in the study of human disease, as several companion papers explore in the context of autoimmune disease, Alzheimer's disease, and cancer."
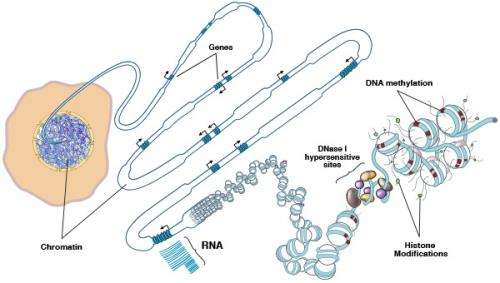
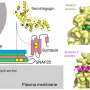
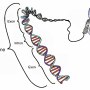
DNA molecules are long, thin double strands containing genes, the discrete units of information that serve as recipes for the protein-making machinery of the cell. In order for DNA molecules to fit into the small space of the cell nucleus, they are compressed and packed like cooked spaghetti, and also wound around spool-like structures called histones. Chemical epigenetic "marks"—the addition of methyl groups in or near genes, and modifications to histones—determine whether genes are available to be transcribed and translated into proteins. Though epigenetic marks are stable, they are reversible, and they can also be altered by environmental factors such as diet, exposure to toxins, and aging. Such changes affect gene expression, which can lead to disease.
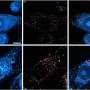
The REMC directed by UCSF's Costello included researchers from UCSF; the University of California, Santa Cruz (UCSC); the University of Southern California (USC), Washington University in St. Louis (WUSTL); and Canada's Michael Smith Genome Sciences Centre and the University of British Columbia (UBC), in Vancouver, Canada. The group provided important data to the REP on several cell types, including epigenomes of the normal human placenta, sperm, breast cells, blood cells, fetal and adult brain cells, and skin cells. Misha Bilenki, PhD, a member of Costello's REMC with an appointment at Canada's Michael Smith Genome Sciences Centre, is co-first author of the Nature paper of which Kellis, of MIT and Harvard, is senior author.
A unique contribution of Costello's REMC was the creation, by WUSTL's Ting Wang, PhD, and David Haussler, PhD, and Jim Kent of UCSC, of the Roadmap Epigenome Browser, a web-based tool that gives scientists worldwide open access to the complete data from the REP.
All told, Costello's REMC contributed four papers, three in Nature Communications and one in Nature Biotechnology, to the 24-paper collection that will appear in print on Feb. 19. In work published online last October in Nature and that will also appear in print as part of the REP collection, UCSF Sandler Faculty Fellow Alex Marson, MD, PhD, and colleagues at The Broad Institute and at Yale School of Medicine showed that epigenomic effects on immune cells can contribute to autoimmune disease.
More information: The research will be freely available to all, and the papers from Nature, along with papers from Nature Communications, Nature Biotechnology, Nature Methods, Nature Neuroscience, Nature Immunology and Nature Protocols, can be explored on the Epigenome Roadmap Site at www.nature.com/epigenomeroadmap