Tracking defects caused by brain tumor mutation yields insight to advance targeted therapy
St. Jude Children's Research Hospital scientists have gained ground toward developing more targeted therapies for the most common childhood brain tumor. The findings appear today in the Journal of Molecular Biology.
The findings involve the DDX3X gene. In 2012, the St. Jude Children's Research Hospital—Washington University Pediatric Cancer Genome Project highlighted DDX3X as a promising focus for efforts to develop targeted therapies against medulloblastoma. Such treatments target the genetic mistakes that give rise to the brain tumor's four subtypes.
The Pediatric Cancer Genome Project found that DDX3X was mutated in a high percentage of patients with the wingless (WNT) subtype of medulloblastoma. Researchers identified eight DDX3X mutations associated with the WNT subtype, which is named for the pathway that is disrupted in the tumors.
Now scientists have determined the DDX3X mutations lead to different molecular defects.
"Our research indicates that mutations of DDX3X do not all lead to a common defect in cells," said Eric Enemark, Ph.D., an associate member of the St. Jude Department of Structural Biology. He and Janet Partridge, Ph.D., associate member of the St. Jude Department of Pathology, are the study's co-corresponding authors. "As a result, therapies that target DDX3X mutations will need to be tailored to account for the specific molecular defects caused by the different disease-associated mutations," Enemark said.
Medulloblastoma is diagnosed in about 400 children and adolescents annually in the U.S., making it the nation's most common pediatric brain tumor. Historically, medulloblastoma has been treated as a single disease even though researchers at St. Jude and elsewhere showed that patient outcomes vary widely based on their tumor subtype. About 11 percent of medulloblastoma patients have the WNT subtype. Nearly all survive with the current treatments that involve surgery, radiation and chemotherapy, but have serious treatment-related side effects.
In this study, scientists used a variety of biochemical, genetic and structural techniques to identify the molecular defects associated with two of the DDX3X mutations associated with medulloblastoma. Investigators demonstrated the mutations interfere with the ability of the DDX3X protein to bind to RNA. Cells use RNA to translate instructions carried by DNA to make the proteins that do the work of cells.
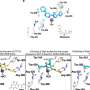
To function properly, DDX3X must bind RNA and also release the chemical energy stored in ATP molecules. In this study, researchers discovered the process depends on a short, flexible loop in DDX3X they call the ATP-binding loop. Without the loop, researchers found that DDX3X and related enzymes could not access energy stored in ATP.
Researchers showed that two DDX3X mutant proteins were unable to tap ATP energy. In contrast, the other DDX3X mutations associated with medulloblastoma exhibited little or no decrease in ATP consumption activity.
Researchers used a yeast model to better understand the consequences of the mutations. If either of the same two mutant DDX3X proteins were substituted for the most similar protein in fission yeast, the yeast died. The yeast survived, however, when the substitution involved the normal human DDX3X protein or DDX3X proteins with other medulloblastoma-associated mutations."Putting the mutant proteins into the yeast model identified which defects were harmful and provided insight into the DDX3X protein function," Partridge said. "We know from previous studies that the fission yeast version of DDX3X is thought to play a role in translation of key regulatory proteins, possibly by helping untangle parts of the RNA molecule." Those proteins include cyclins that regulate cell division, a process often disrupted in cancer.
Enemark added: "We have identified a set of defects for one set of DDX3X mutants that provides a foundation for efforts to develop more individualized treatments for patients with those defects. We still need to determine the molecular defects for the second class of mutants."