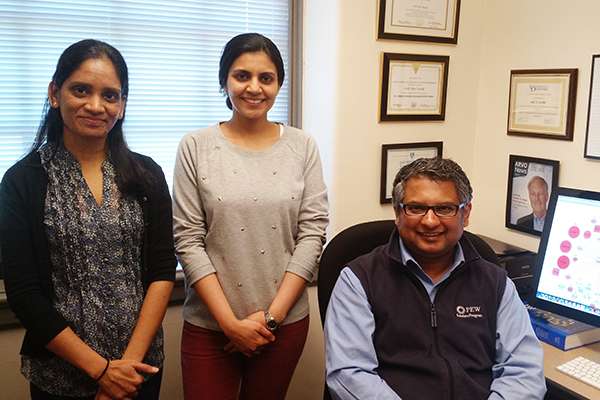
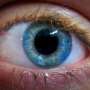
Graduate student Archana Siddam and post-doctoral fellow Deepti Anand studied genetic links to cataracts with Salil Lachke, assistant professor and Pew Scholar .
When cataracts encroach on the eyes, the only effective remedy is to surgically replace the eyes' lenses with synthetic substitutes.
But what if scientists found a way to delay or prevent cataracts from forming in the first place?
Researchers at the University of Delaware may have found such an opportunity by identifying the prime suspects in the formation of cataracts – deficiency of two genes that encode regulatory proteins.
When those two genes are unable to do their work, the lenses of the eyes become cloudy and develop cataracts, no aging process or damaging exposure to radiation required.
Cataracts, the leading cause of blindness, can have a genetic basis.
The discoveries emerged in the laboratory of UD biologist Salil Lachke, assistant professor of biological sciences and a Pew Scholar in biomedical sciences.
Lachke and graduate students Smriti Agrawal, Archana Siddam and post-doctoral fellow Deepti Anand study lens development in mice to better understand the genetic mechanisms that lead to cataracts in humans.
Their findings, published in the journal Human Genetics, could contribute to interventions that one day delay or prevent cataract formation, which now afflicts more than half of the U.S. population over 80 years old and costs Medicare an estimated $3 billion in treatment every year.
The study was supported by a grant from the National Institutes of Health (NIH) and a Fight For Sight grant-in-aid award to Lachke.
Lachke compares the lens of the eye to the lens of a camera. A clear lens, free of internal scratches and other damage, transmits a clear image to the retina. Vision is impaired and can be lost completely when lens deterioration is unaddressed.
Lachke and his students wanted to understand the genetic factors involved in keeping the lens transparent and whether disrupting their function allowed cataracts to form.
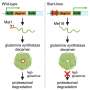
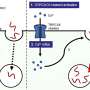
They found answers in the proteins that regulate expression of genes necessary for transparency, demonstrating that deficiency of two regulatory proteins – called Mafg and Mafk – led to formation of cataracts.
The lens of the eye is made up of several kinds of proteins, Lachke said. Some are essential to transcribing genetic information and promoting healthy development.
Smriti Agrawal, shown here at Baylor College of Medicine, was part of the cataract research team in Salil Lachke's lab at the University of Delaware.
To zero in on the genes essential to lens transparency – distinguishing them from normal day-to-day genetic function – Lachke's lab used the integrated Systems Tool for Eye Gene Discovery (iSyTE), a web-based bioinformatics tool initiated by Lachke during his post-doctoral work at Harvard Medical School and now hosted at UD's Center for Bioinformatics and Computational Biology.
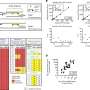
With iSyTE, Lachke and his fellow researchers were able to identify the genes critical to lens formation and develop a detailed roadmap of the network controlled by the Mafg and Mafk proteins.
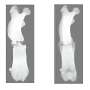
When these regulatory proteins were compromised, several genes responsible for lens transparency were "turned down" and cataracts formed about four months after birth.
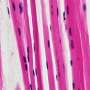
"The lens pathology in the compound mutants is severe," said Agrawal, now studying human genetics in a doctoral program at the Baylor College of Medicine in Texas. "The lens appears to be completely opaque and the fiber cells that comprise the tissue lose their structural organization."
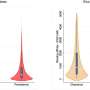
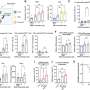
Using a microarray – a collection of thousands of DNA probes on a small chip – researchers can look at the expression of all the genes in the lens, Agrawal said, and bioinformatics allows them to identify the specific targets of regulatory proteins such as Mafg and Mafk.
The discoveries were exciting to be part of, she said.
"When I first found out that the mutant mice exhibited cataract, I ran to Dr. Lachke," she said.
It was a "eureka!" moment, she said.
The iSyTE tool has many other uses, Lachke said, and it is available to other researchers and to the public.
"In just four years, application of the iSyTE tool has significantly expedited eye disease gene discovery, having led to identification of several new – and often unexpected – genes linked to cataracts," he said.
The list of iSyTE-identified genes includes a novel post-transcriptional regulator called TDRD7, commonly involved in germ cell development across various animal species, he said. TDRD7 was shown by Lachke to be linked to human cataract in a research article published in Science.
"There are 22,000 protein-coding genes in our genome - and far less than half are characterized," Lachke said. "Extending the iSyTE approach to other components of the eye and other tissues or organs will greatly expedite disease gene discovery and advance our understanding of the human genome."
Working with Lachke has been an outstanding experience, his three co-authors said.
"He gives us intellectual freedom," Siddam said. "We always go back and run our ideas through him, but he will challenge you and make you think outside the box."
"He likes to include state-of-the-art approaches," Anand said. "He never says 'No' – he tries to understand our ideas."
"And even if they are almost technically impossible, he would never just say 'No,'" Siddam said. "He would appreciate the thought process."
Innovation and discovery happens more freely in such an environment, they said.
"If you want to make breakthroughs, you need to take risks," Anand said.
Agrawal said working with Lachke on cataract research at UD inspired her to pursue her doctorate research on understanding the genetic basis of other eye diseases. Her research now focuses on retinal diseases.
Siddam said top researchers from around the world approached them at a recent conference to discuss the work of the Lachke Lab.
"The credibility of the research here drives us to do more," she said.
More information: "Compound mouse mutants of bZIP transcription factors Mafg and Mafk reveal a regulatory network of non-crystallin genes associated with cataract" Human Genetics July 2015, Volume 134, Issue 7, pp 717-735 DOI: 10.1007/s00439-015-1554-5
Journal information: Human Genetics , Science
Provided by University of Delaware