CanDL database shines light on clinically important cancer gene mutations
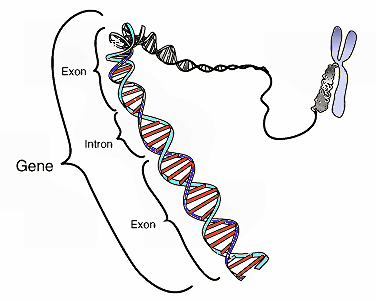
Many clinical trials use genome sequencing to learn which gene mutations are present in a patient's tumor cells. The question is important because targeting the right mutations with the right drugs can stop cancer in its tracks. But it can be difficult to determine whether there is evidence in the medical literature that particular mutations might drive cancer growth and could be targeted by therapy, and which mutations are of no consequence.
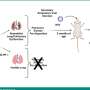
To help molecular pathologists, laboratory directors, bioinformaticians and oncologists identify key mutations that drive tumor growth in tissues obtained during clinical studies, researchers at The Ohio State University Comprehensive Cancer Center - Arthur G. James Cancer Hospital and Richard J. Solove Research Institute (OSUCCC - James) have designed an online database called the Cancer Driver Log, or CanDL.
The freely accessible database is described in a paper published in the Journal of Molecular Diagnostics. It includes mutations in 60 genes, with 334 distinct variants and 169 unique matching literature references across multiple cancers, as of Aug. 24, 2015.
"CanDL is a database of gene mutations that have been functionally characterized or have been targeted clinically or preclinically with approved or investigational agents," says principal investigator and cancer genomics specialist Sameek Roychowdhury, MD, PhD, assistant professor of internal medicine and of pharmacology, and a member of the OSUCCC - James Translational Therapeutics Program.
Entries can be searched by gene or by amino acid variants, or they can be downloaded for custom analyses. The database will be updated quarterly and includes a mechanism for users to contribute novel driver mutations in open collaboration with the Roychowdhury laboratory.
"Currently, pathology laboratories that sequence tumor tissue must manually research the scientific literature for individual mutations to determine whether they are considered a driver or a passenger to facilitate clinical interpretation," Roychowdhury adds. "CanDL expedites this time-consuming process by placing key information about known and possible driver mutations that might be effective targets for drug development at their fingertips," he says.
"CanDL does not tell doctors what to do—it places the evidence in the scientific literature at their fingertips, enabling them to read and interpret the information themselves."
Identifying important driver mutations in a patient's tumor cells can also help reveal why some patients in a clinical trial respond well to a novel agent while others do not respond at all. That information can help improve the effectiveness of existing anticancer drugs, and it can identify subsets of patients who would benefit most from particular therapies.
"Overall, this freely available database will facilitate rapid annotation of cancer genomic testing in molecular pathology labs for mutations," Roychowdhury says.
Other researchers from Ohio State involved in this study were Senthilkumar Damodaran, Jharna Miya, Esko Kautto, Eliot Zhu, Eric Samorodnitsky, Jharna Datta and Julie W. Reeser.