Predictive model could help guide choices for breast cancer therapies

Biomedical engineers have demonstrated a proof-of-principle technique that could give women and their oncologists more personalized information to help them choose options for treating breast cancer.
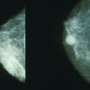
Thanks to diagnostic tests, clinicians and patients can already know the type of breast cancer they're up against, but one big question remains: How likely is it that the cancer will invade other parts of the body? Answering that question could help guide the choice of treatment options, from aggressive and difficult therapies to more conservative ones.
By studying chemical signals from specific cells that are involved in helping cancer invade other tissues in each woman's body, researchers have developed a predictive model that could provide an invasiveness index for each patient.
"We want women to have more information to make a personal decision beyond the averages calculated for an entire population," said Manu Platt, an associate professor in the Department of Biomedical Engineering at Georgia Tech and Emory University. "We are using our systems biology tools and predictive medicine approaches to look at potential markers we could use to help us understand the risk each woman has. This would provide information for a more educated discussion of treatment options."
The research, sponsored with funds from the Georgia Research Alliance and the Giglio family donation to the Department of Biomedical Engineering, was reported September 9 in the journal Scientific Reports. Beyond breast cancer, the technique could offer similar decision-making assistance for men with prostate cancer, where treatment also requires making difficult choices about the risk of metastasis.
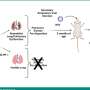
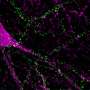
Platt's research team is examining chemical signals produced by the macrophages that can help aggressive tumors invade new tissues. Macrophages normally clean up foreign particles and harmful microorganisms in the body, but aggressive tumors can enlist macrophages in helping them metastasize. Tumor associated macrophages contribute significantly to tumor invasion, with cysteine cathepsin proteases - enzymes that break down proteins in the body - important contributors.

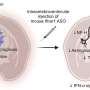
To develop their predictive index, Platt's research team used variability in macrophage expression of four types of cathepsin, the cathepsin inhibitor cystatin C, and kinase activation levels. The model, which has been under development for two years, was produced by studying macrophages from a population of women who didn't have breast cancer. Platt and colleagues Keon-Young Park and Gande Li co-cultured a standard breast cancer cell line (MCF-7) with macrophages produced from monocytes donated by these cancer-free women.
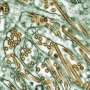
Next, they measured the level of invasiveness facilitated by macrophages from each individual donor, exposing the cancer cells and macrophages to a collagen gel designed to simulate breast tissue and measuring how many cells invaded it. While the breast is composed of many other tissues, collagen makes up the largest proportion and provided a good measure of how aggressively the cells would invade, Platt said.
Platt's team correlated the level of invasion through the gel to the chemical signals being expressed by the macrophages. The researchers were surprised at the large amount of patient-to-patient variability in macrophage activity - variability that could account for the outcome differences in the patients receiving similar cancer treatments. The signaling levels and related invasion measurements were used to train a computational model developed by Platt's team.
The researchers next obtained blood samples containing monocytes from nine patients being treated for breast cancer at DeKalb Medical Center, a major Atlanta-area hospital. They measured signals from these macrophages and used their model - which had been trained on macrophage signaling and resulting invasiveness - to predict which of the cancer patients would be expected to have more invasive types of cancer. They compared their predications to what the clinician - Dr. John Kennedy - provided as their initial diagnosis.
"Based on the cells we got from the clinic, the ones that had been predicted to have the greatest potential for invasion were the ones that had produced the most invasive form of breast cancer in the patients," Platt said.
While the study could not account for possible differences in the length of time the cancers had been growing, they did correlate well with observations. In future research, Platt hopes to follow the women for five years to determine if the model's predictions are related to cancer recurrence. He also plans to expand the model with additional macrophage data, and test it against additional blood samples.
"The more information you give the model, the closer you get to the prediction," he said. "We think this is a very big start."
The strength of this technique, Platt believes, is that it measures what's happening at the level where cancer is metastasizing.
"We are measuring at the level of activity of these intracellular enzymes and the ultimate activity of the proteases they produce that are not only the biomarkers of the tumor, but also help the tumor grow," he said. "Everything about us is different. Our genetics are different and our lifestyles are different, so clinicians have to make decisions in all that variability. All of those differences can be measured and captured in this output."
Platt believes the technique could one day lead to a simple blood test that would provide information useful in making therapy recommendations. The test might also help determine which women should be monitored more closely to detect the beginnings of a cancer.
"Together, this establishes proof-of-principle that personalized information acquired from minimally invasive blood draws may provide useful information to inform oncologists and patients of invasive/metastatic risk, helping to make decisions regard radical mastectomy or milder, conservative treatments to save patients from hardship and surgical recovery," he wrote in the paper.