Research connects specific variations in RNA splicing with breast cancer causation
Researchers have identified cellular changes that may play a role in converting normal breast cells into tumors. Targeting these changes could potentially lead to therapies for some forms of breast cancer.
In work published today in Molecular Cell, a team of researchers led by Professor Adrian Krainer at Cold Spring Harbor Laboratory (CSHL) analyzed the effects of overexpressing a cancer-promoting protein called SRSF1 in both a cellular model of breast cancer and in human breast tumors, and identified changes that could be responsible for SRSF1's ability to cause cancer.
SRSF1 is a splicing factor, a type of protein that plays a crucial role in the process by which genes give rise to proteins. Splicing factors such as SRSF1 help edit the coded "message" copied from genes. These copies of the gene's message, called transcripts, are made of RNA. Splicing literally snips out parts of the message - called introns - that don't encode protein; and they paste together the remaining portions of the message - called exons - that do encode protein.
SRSF1 and other splicing factors can also direct a related process called alternative splicing—the mixing and matching of different exons from one gene, which, when spliced together, give rise to variant versions of a single protein, ones that often have very different functional properties.
"Long ago we and others saw that the levels of splicing proteins are altered in various cancers," says Krainer. "Previous work indicated that there are lots of alterations in splicing in the context of cancer," he says.
SRSF1 was previously shown to be an oncoprotein, i.e., a protein involved in cancer. When it is overexpressed, it can help convert normal cells into tumor cells. Krainer decided to examine its role in breast cancer "because we see frequent overexpression of SRSF1 in breast tumors," he says.
Experiments conducted by Olga Anczuków and Martin Akerman, co-first authors of the new paper, other colleagues in the Krainer lab, and their collaborators at the ETH in Zurich, demonstrated how overexpression of SRSF1 in a cellular model of breast cancer resulted in changes in the RNA messages generated in the cells. This allowed the team to pinpoint the splicing targets regulated by SRSF1, and gave them a glimpse of the SRSF1-related changes in those cells.
The team identified and validated hundreds of alternative splicing events regulated by SRSF1. They also looked at data from The Cancer Genome Atlas, identifying alternative splicing events associated with SRSF1 overexpression in human breast tumors. "We pay special attention to potential targets that overlap between all these different data sets, because it's a way of narrowing down or giving higher priority to targets that may play an important role," explains Anczuków.
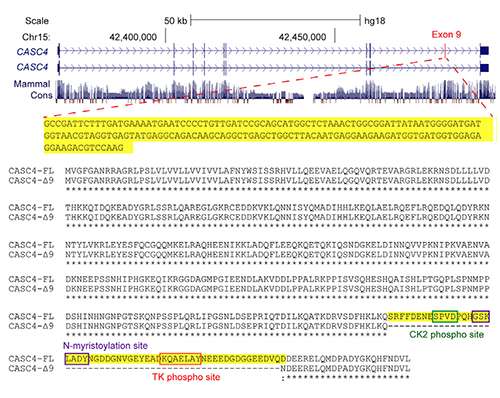
Based on the overlap between SRSF1 overexpression data in breast cancer cell lines and human breast tumors, the researchers identified a candidate gene called CASC4. Overexpressing one alternatively-spliced form of the CASC4 protein partially mimicked the effects of overexpressing SRSF1, suggesting that CASC4 contributes to SRSF1's oncogenic effects. Krainer says CASC4 is likely to be just one of many targets that explain SRSF1's ability to cause cancer, and several targets probably act in concert to produce all the oncogenic changes related to SRSF1 overexpression.
Identifying the key oncogenic alternative splicing events caused by SRSF1 and other splicing factors could lead to potential therapeutic targets for treating breast cancer. "If we find that a particular change is really critical in tumor maintenance, then we could potentially develop therapies to restore the normal splicing pattern," says Krainer. "It probably depends on there being a small number, ideally one, but maybe a couple of crucial targets that the tumors really depend upon."
More information: "SRSF1-Regulated Alternative Splicing in Breast Cancer" appears online in Molecular Cell on October 1, 2015. The authors are: Olga Anczuków, Martin Akerman, Antoine Cléry, ..., Yimin Hua, Frédéric H.-T. Allain, Adrian R. Krainer. The paper can be obtained online at: www.sciencedirect.com/science/journal/aip/10972765