One course of antibiotics can affect diversity of microorganisms in the gut
A single course of antibiotics has enough strength to disrupt the normal makeup of microorganisms in the gut for as long as a year, potentially leading to antibiotic resistance, European researchers reported this week in mBio, an online open-access journal of the American Society for Microbiology.
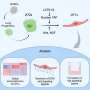
In a study of 66 healthy adults prescribed different antibiotics, the drugs were found to enrich genes associated with antibiotic resistance and to severely affect microbial diversity in the gut for months after exposure. By contrast, microorganisms in the saliva showed signs of recovery in as little as few weeks.
The microorganisms in study participants' feces were severely affected by most antibiotics for months, said lead study author Egija Zaura, PhD, an associate professor in oral microbial ecology at the Academic Centre for Dentistry in Amsterdam, the Netherlands. In particular, researchers saw a decline in the abundance of health-associated species that produce butyrate, a substance that inhibits inflammation, cancer formation and stress in the gut.
"My message would be that antibiotics should only be used when really, really necessary," Zaura said. "Even a single antibiotic treatment in healthy individuals contributes to the risk of resistance development and leads to long-lasting detrimental shifts in the gut microbiome."
It's not clear why the oral cavity returns to normal sooner than the gut, Zaura said, but it could be because the gut is exposed to a longer period of antibiotics. Another possibility, she said, is that the oral cavity is intrinsically more resilient toward stress because it is exposed to different stressors every day.
The investigators enrolled healthy adult volunteers from the United Kingdom and Sweden. Participants were randomly assigned to receive a full course of one of four antibiotics (ciprofloxacin, clindamycin, amoxicillin or minocycline) or a placebo. The researchers, who did not know which medication participants took, collected fecal and saliva samples from the participants at the start of the study; immediately after taking the study drugs; and one, two, four and 12 months after finishing the medications. They performed a laboratory technique called 16S rRNA gene amplicon sequencing, which can identify the presence of bacteria, on 389 fecal and 391 saliva samples. Then, they performed another lab technique called metagenomic shotgun sequencing on samples where researchers saw the largest differences before and after antibiotic usage, to study the emergence of antibiotic resistance.
Researchers found that participants from the United Kingdom started the study with more antibiotic resistance than did the participants from Sweden, which could result from cultural differences. There has been a significant decline in antibiotic use in Sweden over the last two decades, Zaura said.
In addition, fecal microbiome diversity was significantly reduced for up to four months in participants taking clindamycin and up to 12 months in those taking ciprofloxacin, though those drugs only altered the oral cavity microbiome up to one week after drug exposure. Exposure to amoxicillin had no significant effect on microbiome diversity in either the gut or oral cavity but was associated with the greatest number of antibiotic-resistant genes.
"Certainly we cannot live or survive without antibiotics; that's out of the question," she said. "But there are situations when we should not use them, like when there are no evidence-based reasons."
Further study to understand the mechanisms behind the oral microbiome's resilience might prove useful in combating microbial imbalances in the gut, she said.
More information: E. Zaura et al. Same Exposure but Two Radically Different Responses to Antibiotics: Resilience of the Salivary Microbiome versus Long-Term Microbial Shifts in Feces, mBio (2015). DOI: 10.1128/mBio.01693-15